How a tension sensor plays integral role in aligned chromosome partitioning
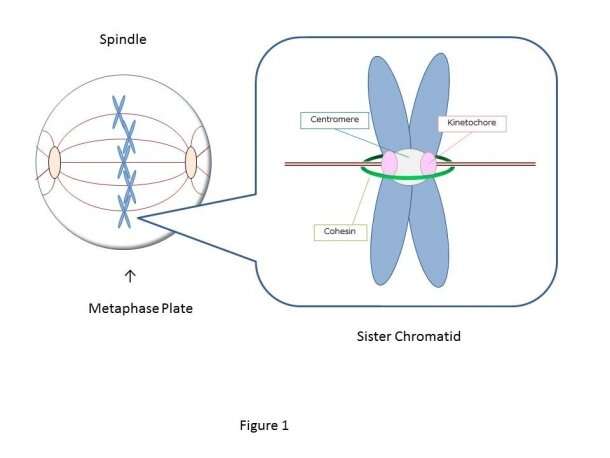
A Waseda University-led research uncovered the molecular mechanism of how a particular cancer-causing oncogene could trigger an onset of acute myeloid leukemia (AML).
Characterized by symptoms such as fatigue, shortness of breath and bleeding gums, AML is a type of cancer that starts in the bone marrow and quickly affects blood due to rapid growth of leukemia cells. This abnormality is caused by mutated genes in chromosomes, turning on oncogenes while turning off tumor-suppressing genes. Mutation occurs when chromosomes are not properly replicated during cell division, and misalignment of even one out the total of 46 chromosomes in a cell causes missegregation.
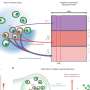
To prevent such chromosomal abnormalities, a cell precisely controls chromosome distribution to the new-born cells with its tension sensor, which locates the central region of the duplicated chromosomes called the centromere and detects whether the attached microtubule is applying the right among of force to the kinetochore, a protein complex at the centromere.
"Since discovering the enzyme Aurora B kinase (Aurora B) in 1996, we have found that Aurora B plays an integral role as a tension sensor by adjusting microtubule attachment to the kinetochore for chromosome regulation, and that protein phosphatase 2 (PP2A) also acts as a tension sensor by controlling chromosome alignment in correlation with Aurora B," said Professor Yasuhiko Terada of Waseda University in Tokyo. "However, the system is extremely complex, and its molecular mechanism was not well understood."
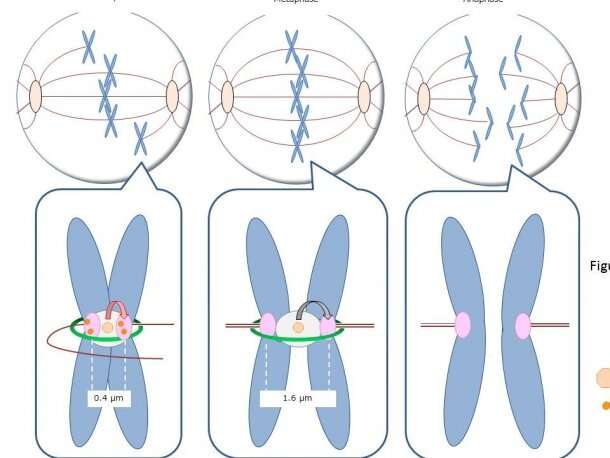
In their latest study published in the Journal of Cell Biology, Terada's team found that SET/TAF1, a proto-oncogene of AML, also functions as a tension sensor by fine-tuning the enzyme activities of Aurora B and PP2A. As the three tension sensors interact with each other, the replicated chromosomes are distributed evenly to the new-born cells and prevent chromosome abnormalities.
"The protein encoded by the SET oncogene (SET) maintained Aurora B activity by inhibiting the PP2A activity at the centromeres. Also, it was intriguing to observe that though SET inhibits activity of PP2A to allow a high Aurora B activity and adjust microtubule attachment when chromosomes are not aligned, it detaches from the kinetochore and decreases the activity of Aurora B to stabilize attachment of microtubules to the kinetochore when chromosomes are aligned," Terada explained.
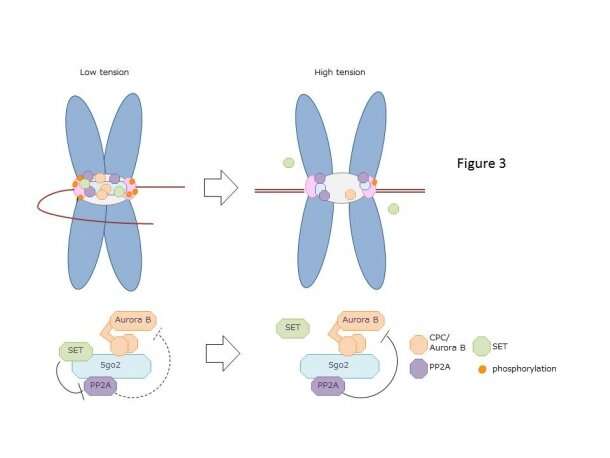
Additionally, experiments using molecular biological techniques investigated the oncogenic function of SET to study whether centromere localization of SET is essential for chromosomal abnormality. Results showed that SET disrupts the tension sensor mechanism at the centromere, supporting previous research that report how abnormal activity of Aurora B is observed in many cancer cells, and how overexpression of Aurora B in normal cells induce chromosome misalignment.
Though many questions remain unanswered to fully understand the molecular mechanism of the tension sensor, Terada believes that this discovery could serve as a baseline for further investigation to elucidate the molecular mechanism of cancer malignancy by chromosome missegregation and the development of leukemia, as well as to create anticancer drugs that target SET and Aurora B.
More information: Yuichiro Asai et al, Aurora B kinase activity is regulated by SET/TAF1 on Sgo2 at the inner centromere, The Journal of Cell Biology (2019). DOI: 10.1083/jcb.201811060
Journal information: Journal of Cell Biology
Provided by Waseda University