Genome mining reveals novel production pathway for promising malaria treatment

Microbes are well-known among biologists as master engineers of useful small molecules, and there are many tricks of their trade. When researchers at the University of Illinois took a closer look at how a known microbe makes a known so-called natural product, they were rewarded with the discovery of a completely unknown biochemical trick.
G. William Arends Professor of Molecular and Cellular Biology at the University of Illinois William Metcalf led the study with then-postdoctoral researcher Elizabeth (Betsy) Parkinson. Parkinson is now an assistant professor of chemistry at Purdue University. Metcalf, Parkinson and coauthors reported their work, which was supported by NIH, in Nature Chemical Biology.
The work began with a surprise: the researchers set out to explore how their microbe of interest, Streptomyces lavendulae, creates a chemical called fosmidomycin. The team was interested in how this compound is created in part because it's an antimicrobial that is effective against malaria, a mosquito-borne illness that kills hundreds of thousands of people each year. As expected, S. lavendulae did produce a compound that killed microbes—but it wasn't fosmidomycin.
"The most interesting stuff research is where you ask a question and you get a completely unexpected answer," Metcalf said. "Something turn out as we expected; that's great!"
More surprises quickly followed. The team traced the bacterium's killing powers to production of a closely related molecule, dehydrofosmidomycin, a known natural product that may even be slightly better than fosmidomycin for treating malaria. However, the genes that S. lavendulae was using to make dehydrofosmidomycin were completely unlike those seen in other microbes.
"It's very similar to another class of molecule that we've worked on in the past, virtually identically, chemically and structurally, but the biosynthetic pathway and the genes are completely different," Metcalf said. "Which if you think about evolution and how you got there, that's fascinating, that these molecules are so good that nature independently discovered it multiple times."
Microbes evolve the capability to make natural products like fosmidomycin and dehydrofosmidomycin to help them outcompete neighboring microbes for space and resources. Each natural product is chemically crafted by a series of proteins called enzymes, which take turns tweaking the growing molecule by adding or removing atoms to change its shape and activity. Microbial genomes are scattered with clusters of genes encoding these enzymes, with one cluster typically containing all the genes necessary for making one natural product.
Metcalf's laboratory and other researchers at the Carl R. Woese Institute for Genomic Biology at the University of Illinois want to explore the relationship between microbial natural products and the gene clusters that enable their production. By learning to recognize what genes lead to what types of products, they hope to use genome sequencing to speed discovery of new natural products that, like fosmidomycin and related molecules, may have key therapeutic properties.
Metcalf was particularly excited to see a familiar type of molecule being made by an unfamiliar gene cluster.
"The technical term is convergent evolution towards a chemical product," Metcalf said. "And that tells you . . . that it's a really good molecule. It does what nature wants it to do: it's an antibacterial and it also kills parasites, like malaria and plants, like weeds, it's really got a lot of uses. It's utterly non-toxic to human beings, which is nice."
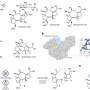
The researchers delved deeper into the details of the new gene cluster and the chemical reactions facilitated by its enzymes. They reconstructed and experimentally confirmed a series of steps leading from the starting "ingredients" to the finished product.
"So why do you care about how molecules like this are made? . . . A really good bioengineered pathway, it's the cheapest way to make anything," Metcalf said. "This offers another route to the same molecule, which might be a more efficient route, might be a cheaper route, that has yet to be explored."
The highlight of the newly discovered pathway was an enzyme encoded by the gene dfmD. Its name, reminiscent of a library call number and chosen by the researchers to indicate its position in the dehydrofosmidomycin-producing gene cluster, belies the novelty of the chemical reaction the enzyme facilitates.
"You break two carbon-nitrogen bonds, you reform one carbon-carbon bond, and you oxidize another carbon-carbon bond. And you do that all in one step," Metcalf said. In other words, the enzyme breaks a piece off the larger molecule, flips it around, reattaches it, and tweaks the resulting product, all in the single continuous action, analogous to a person changing seat configurations in a minivan commercial.
"In simplest terms, what dfmD is doing is a chemical reaction that's not easy to envision, number one, just based on first principles of chemistry; and number two, that's never been observed in nature before," Metcalf said. "Because this is doing something radically different, it adds to that body of knowledge so that when we look at new pathways, we can think about how they might work."
More information: Elizabeth I. Parkinson et al, Fosmidomycin biosynthesis diverges from related phosphonate natural products, Nature Chemical Biology (2019). DOI: 10.1038/s41589-019-0343-1
Journal information: Nature Chemical Biology
Provided by University of Illinois at Urbana-Champaign