New method increases accuracy of nontuberculous mycobacteria identification
The bacterial genus Mycobacterium has the dubious honor of including species responsible for two of the best-known chronic human infectious diseases: tuberculosis and leprosy. But unlike their more famous cousins, for which effective treatment strategies have long been available, it is the 200 or so lesser known Mycobacterium species that are currently causing a resurgence in pulmonary diseases in recent times.
Referred to collectively as nontuberculous mycobacteria (NTM), these species are widely found in soil and water. However, if given the chance, NTM can cause serious skin and lung infections in susceptible patients. One of the biggest impediments to the treatment of NTM infections is the difficulty in telling the bacteria apart, particularly at the subspecies level. Accurate identification is crucial though, as different species show varying levels of responsiveness to different antibiotic therapies.
To address the lack of an accurate and sensitive identification method for NTM, a research team from Osaka University and the University of the Ryukyus in Japan have developed software that reliably identifies NTM based on sequence data from bacterial genes. In a paper published this month in Emerging Microbes and Infections, the researchers explain how they developed the software and what it means for the treatment of NTM infections.
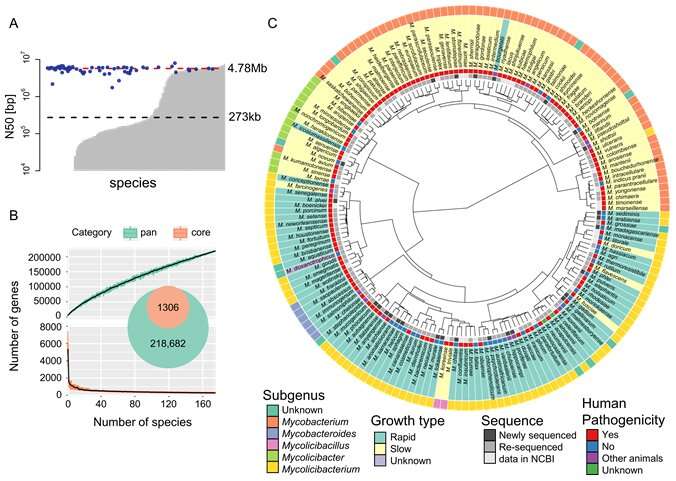
"Among the current NTM identification methods, the most sensitive and accurate are based on genomic information," says author Shota Nakamura. "However, despite the recognized need for high quality genomic data that allows NTM identification to the subspecies level, current databases only contain assemblies for 148 species. Therefore, in this study, we sequenced the genomes of a further 27 species and resequenced the genomes of 36 species."
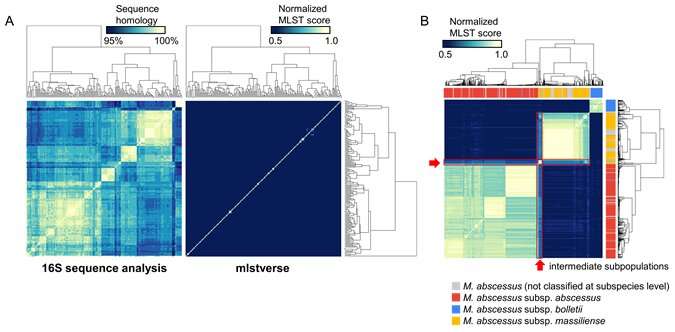
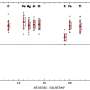
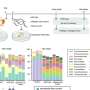
Using their newly acquired sequences in conjunction with 7,484 previously published genome assemblies, the researchers developed a comprehensive database of 175 NTM species identified based on the sequences of 184 separate genes. Known as multilocus sequence typing, each species can be differentiated by its specific combination of sequence differences within the 184 genes. The database also included other Mycobacterium species for comparison.
"Once we had assembled our database, we developed software, called mlstverse, that compares unknown sequences against the database, resulting in accurate identification of NTM," explains lead author of the study Yuki Matsumoto. "When we compared our method with other approaches for the identification of 29 clinical NTM isolates, mlstverse was the only method that identified all 29 isolates to the subspecies level."
The possible applications are promising—Takeshi Kinjo, a medical doctor in the University of Ryukyus Hospital, says that "this method can potentially be used to identify NTM from clinical specimens, allowing the implementation of targeted therapies and improving the cure rates of NTM-associated infections."
More information: Yuki Matsumoto et al. Comprehensive subspecies identification of 175 nontuberculous mycobacteria species based on 7547 genomic profiles, Emerging Microbes & Infections (2019). DOI: 10.1080/22221751.2019.1637702
Provided by Osaka University