Engineered 'smart cells' improve production of pharmaceutical raw materials
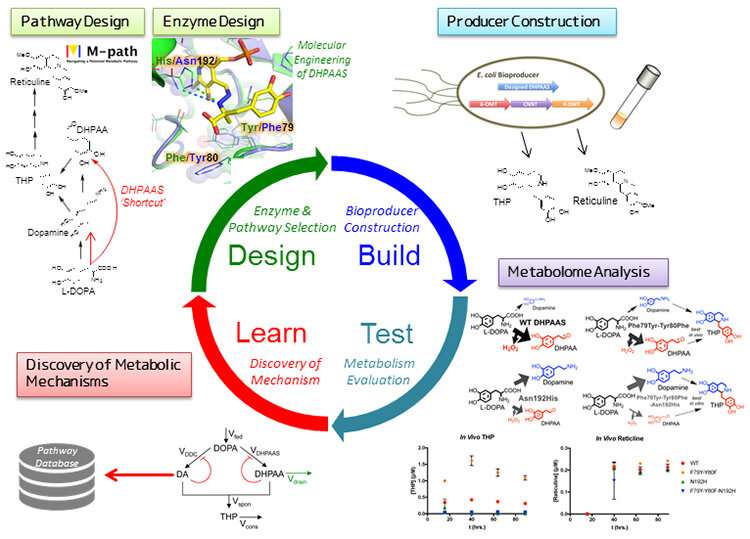
Researchers in Japan have developed an integrated synthetic biology system to construct new metabolic pathways and enzymes within microbes. By incorporating a "Design, Build, Test, Learn" (DBTL) workflow, the production of pharmaceutical raw materials could be systematically optimized. This application supports the concept of the DBTL workflow as a sustainable method for production of complex and valuable materials. The results were published on May 1 in the open access journal Nature Communications.
This study is part of a project from the New Energy and Industrial Technology Development Organization (NEDO), and was carried out by a Kobe University research team led by Assistant Professor Christopher Vavricka, Visiting Professor Michihiro Araki, Professor Tomohisa Hasunuma and Professor Akihiko Kondo. Close collaboration with a research team led by Associate Professor Hiromichi Minami (Research Institute for Bioresources and Biotechnology, Ishikawa Prefectural University) was also central to this project.
The collaborative research team is participating in a NEDO research and development project under the theme of "Development of Production Techniques for Highly Functional Biomaterials Using Smart Cells of Plants and Other Organisms (Smart Cell Project)". The goal of the Smart Cell Project is to achieve mass production of highly valued target materials by introducing genes that encode improved pathways into host microbes. This process relies heavily upon information analysis technology to re-design metabolic systems and pathways that can increase production amounts and production efficiency.
Alkaloid production was selected as a prime example for optimization because alkaloids are key intermediates in the production of pharmaceuticals including opioid pain medications. Recently, the production of alkaloid derived pain medicine has been achieved using microbes, but to make this commercially viable production yields must be improved. The key alkaloid intermediate tetrahydropapaveroline (THP) was previously produced using a combination of two enzymes: aromatic L-amino acid decarboxylase (AAAD) and monoamine oxidase (MAO). However, the relaxed specificity MAO has been a barrier to efficient THP production.
To improve this process, a metabolic design program called M-path was put to the test. This prediction software was developed by Professor Araki at Kobe University, and applied to identify novel enzymes that can bypass MAO for improved pathways to the key alkaloid intermediate THP. The M-path analysis led to the discovery of a promising natural enzyme found in silkworms called 3,4-dihydoxyphenylacetaldehyde synthase (DHPAAS) as an alternative to MAO. DHPAAS is novel in that it possesses amine oxidizing ability in additional to conventional decarboxylation activity. The team then developed structure-based enzyme engineering methods to identify key amino acids involved in determining DHPAAS enzyme activity. This enabled them to create artificial DHPAAS enzymes that can tune the ratio of decarboxylase and amine oxidase activities, leading to improved production of the key intermediate THP.
When the team introduced the newly designed metabolic pathway, including engineered enzymes, into the conventional laboratory bacteriumEscherichia coli, they were able to precisely control the ratio of key intermediates dopamine (decarboxylation production) and DHPAA (oxidation product). Balancing dopamine and DHPAA levels led to improved alkaloid production within the re-designed "smart cells". To optimize the microbial production system further, over 100 metabolites were analyzed with Shimadzu mass analysis systems, enabling the team to identify bottleneck reactions and by-product forming side reactions. By incorporating the metabolite information as learning data to drive forward a new DBTL cycle, production of downstream alkaloid intermediates was further improved.
These findings demonstrate that combining advanced biotechnology and computer science is an effective strategy to rapidly develop cell factories that can produce many different kinds of valuable materials. In addition, the ability to engineer artificial enzyme functions can help to expand the range of possible production targets. Looking forward, the authors believe that the DBTL workflow will enable more efficient production of various useful materials, including pharmaceuticals, fine chemicals, biological chemicals and biofuels. This synthetic biology workflow is expected to make significant contributions to the next-generation smart cell industry for the production of complex pharmaceuticals and chemicals as well as newly discovered materials.
More information: Christopher J. Vavricka et al. Mechanism-based tuning of insect 3,4-dihydroxyphenylacetaldehyde synthase for synthetic bioproduction of benzylisoquinoline alkaloids, Nature Communications (2019). DOI: 10.1038/s41467-019-09610-2
Journal information: Nature Communications
Provided by Kobe University