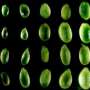
Seed abortion and the role of RNA Pol IV in seed development

ASPB is pleased to announce the publication in The Plant Cell of important research that explores the cause of seed abortion and the role of the enzyme RNA Pol IV to understand how seed development works.
In flowering plants, the embryo is surrounded by the endosperm. Endosperm tissue mediates nutrient transfer between the growing embryo and the mother. The endosperm is distinct from the rest of the plant because it has only one copy of the father's genome and two copies of the mother's. The ratio of maternal to paternal genomes is remarkable because of its importance to seed viability and development. Seeds with extra genomes that alter this critical ratio undergo a process known as interploidy seed abortion due to defective endosperm development. RNA Pol IV is an enzyme specific to plant genomes that generates small RNA molecules that silence gene expression from transposons and repetitive DNA, playing a major role in defending the genome against viruses and transposable elements. The new work shows that RNA Pol IV plays a key role in interploidy seed abortion.
This research, coauthored by P.R.V. Satyaki and Mary Gehring of the Whitehead Institute for Biomedical Research, focused on the following questions: How does the lack of RNA Pol IV prevent interploidy seed abortion? Where does RNA Pol IV act, in the endosperm or in the father, to influence gene expression in the endosperm? In what genetic pathway does RNA Pol IV function to cause seed abortion?
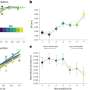
In this article, Satyaki and Gehring demonstrate that RNA Pol IV targets genes in the father via the "canonical" RNA-directed DNA methylation pathway, a major gene-silencing pathway in Arabidopsis plants, resulting in interploidy seed abortion. The researchers compared gene transcription in the endosperm of aborted interploidy seeds with that of seeds that were viable due to the loss of paternal RNA Pol IV. The researchers found that transposons and thousands of genes, even imprinted ones, were misregulated in both living and dying seeds. The researchers learned that misregulation of a relatively small number of genes sets living seeds apart from aborting ones.
This study is also important because it identified a transcriptional buffering system in the endosperm. This system counteracts the effects of a higher dose of the paternal genome by reducing the expression of the paternal copies of some genes and increasing the expression of maternal copies of other genes.
First author P.R.V. Satyaki said: "The next steps are to unravel the mechanism underlying the transcriptional buffering system and to identify the genes responsible for interploidy seed abortion using the shortlist of candidate genes generated from our transcriptional studies."
More information: Satyaki, P.R.V., and Gering, M. (2019). Paternally-acting canonical RNA-directed DNA methylation pathway genes sensitize Arabidopsis endosperm to paternal genome dosage. Plant Cell doi: 10.1105/tpc.19.00047
Journal information: Plant Cell
Provided by American Society of Plant Biologists