Researchers characterize molecular scissors for plastic waste
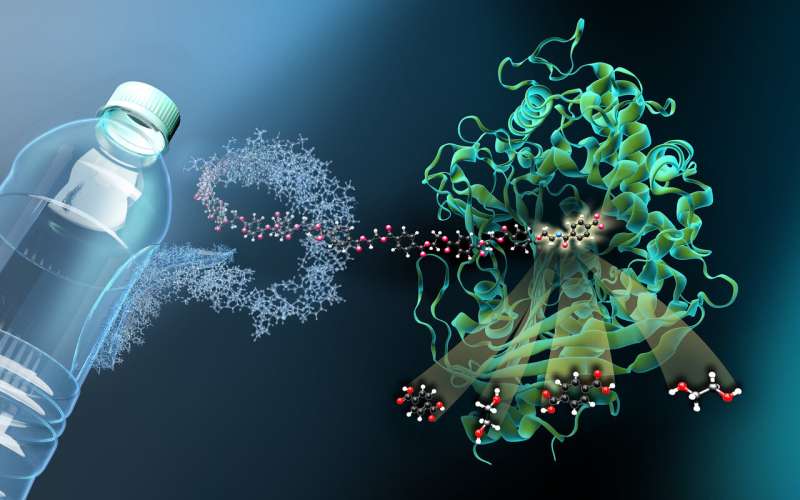
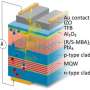
A research team from the University of Greifswald and Helmholtz-Zentrum-Berlin (HZB) has solved the molecular structure of the enzyme MHETase at BESSY II. MHETase was discovered in bacteria, and together with a second enzyme, PETase, is able to break down the widely used plastic PET into its basic building blocks. This 3D structure already allowed the researchers to produce a MHETase variant with optimized activity in order to use it, together with PETase, for a sustainable recycling of PET.
Plastics are extremely versatile and almost eternally durable. But this is also a problem, because after only about 100 years of producing plastics, plastic particles are now found everywhere—in groundwater, in the oceans, in the air, and in the food chain. Around 50 million tonnes of the industrially important polymer PET are produced every year. Just a tiny fraction of plastic is currently recycled via expensive and energy-consuming processes that yield either downgraded products or depend in turn on adding 'fresh' crude oil.
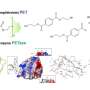
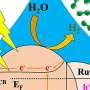
In 2016, a group of Japanese researchers discovered a bacterium that grows on PET and partially feeds on it. They found that the bacterium possesses two special enzymes, PETase and MHETase, that digest PET plastic polymers. PETase breaks down the plastic into smaller PET building blocks, primarily MHET, and MHETase splits this into the two basic precursor building blocks of PET, terephthalic acid and ethylene glycol. Both components are valuable for synthesising new PET without the addition of crude oil for a closed sustainable production and recovery cycle.
In April 2018, the structure of PETase was finally solved independently by several research groups. The Diamond Light Source was also involved in the experiments. However, PETase is only part of the solution. It is equally important to characterize the structure of the second enzyme, MHETase.
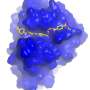
"MHETase is considerably larger than PETase and even more complex. A single MHETase molecule consists of 600 amino acids, or about 4000 atoms. MHETase has a surface that is about twice as large as the surface of PETase and has therefore considerably more potential for optimization toward decomposition of PET," explains biochemist and structural biologist Dr. Gert Weber from Helmholtz-Zentrum Berlin and Freie Universität Berlin.
During an interim professorship at the University of Greifswald, Weber contacted the biotechnologist Prof. Uwe Bornscheuer at the Institute of Biochemistry, who was already involved with plastic-degrading enzymes. Together, they developed the idea of solving the structure of MHETase and then using this insight to optimise the enzyme for applications in PET recycling. To do this, they first had to extract the enzyme from bacterial cells and purify it. Within this collaboration, the teams succeeded in obtaining the complex three-dimensional architecture of MHETase at BESSY II, the synchrotron source at HZB in Berlin.
"In order to see how MHETase binds to PET and decomposes it, you need a fragment of plastic that binds to MHETase but is not cleaved by it," explains Weber. A member of Weber's prior research team in Greifswald, Dr. Gottfried Palm, cut up a PET bottle, chemically decomposed the PET polymer and synthesised a small chemical fragment from it that binds to MHETase but can no longer be cleaved by it. From this 'blocked' MHETase, tiny crystals were grown for structural investigations at the HZB. "The structural investigations enabled us to watch MHETase virtually 'at work' and develop strategies for how to optimise this enzyme," explains Weber.
"Thanks to the joint research group format, we have the means to offer beamtime access on the highly demanded BESSY II MX beamlines for measurements very quickly at any time," says Dr. Manfred Weiss, who is responsible for the BESSY II MX beamlines. The three-dimensional architecture of MHETase actually displays some special features: enzymes such as MHETase bind to their target molecule first before a chemical reaction occurs. For breakdown of a molecule you need a tailor-made enzyme: "We can now exactly localise where the MHET molecule docks to MHETase and how MHET is then split into its two building blocks terephthalic acid and ethylene glycol," says Weber.
However, neither PETase nor MHETase are particularly efficient. "Plastics have only been around on this scale for a few decades—even bacteria with their rapid successions of generations and rapid adaptability have not managed to develop a perfect solution through the evolutionary process of trial and error over such a short time," explains Weber. "Thanks to the clarification of the structure of this very important enzyme, we have now also been able to plan, produce and biochemically characterise variants that show significantly higher activity than natural MHETase and are even active against another intermediate product of PET degradation, BHET," adds Uwe Bornscheuer.
In future, Uwe Bornscheuer will work on systematically optimising the enzymes PETase and MHETase for their task—the decomposition of PET. Gert Weber plans to supplement these studies with further work on biological structures in order to systematically develop plastic-digesting enzymes for environmental applications. Access to the measuring stations and the IT infrastructure of HZB is indispensable for this.
Producing these kinds of enzymes in closed biotechnological cycles, for example, could be a way to really break down PET plastics and other polymers into their basic building blocks. This would also be the key to ideal recycling and a long-term solution to the plastic waste problem: production of plastic would be a closed cycle and no longer dependent on crude oil.
The study is published in Nature Communications.
More information: Nature Communications, DOI: 10.1038/s41467-019-09326-3
Journal information: Nature Communications
Provided by Helmholtz Association of German Research Centres