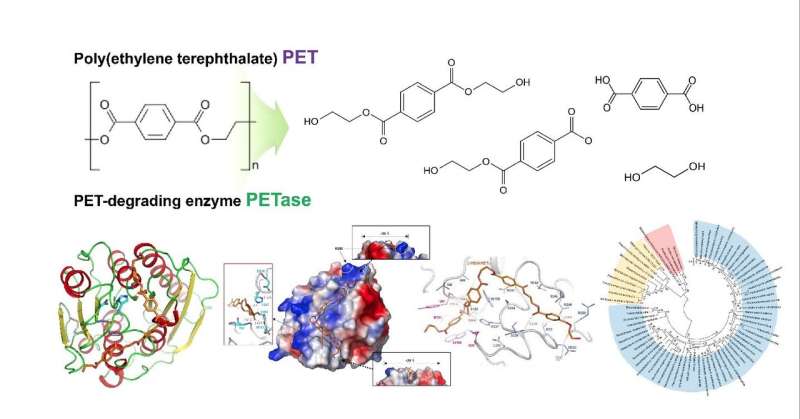
Structural insight into the molecular mechanism of PET degradation
A KAIST metabolic engineering research team has found a molecular mechanism showing superior degradability of polyethylene terephthalate (PET). This is the first report to determine the 3-D crystal structure of Ideonella sakaiensis PETase and develop the new variant with enhanced PET degradation.
Research projects have been working to address the non-degradability of materials. A polyethylene terephthalate (PET)-degrading bacterium called Ideonella sakaiensis was recently identified for the possible degradation and recycling of PET by a Japanese team that published results in Science in 2016. However, the detailed molecular mechanism of PET degradation was not determined.
The team led by Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering and another team team led by Professor Kyung-Jin Kim of the Department of Biotechnology at Kyungpook National University conducted this research. The findings were published in Nature Communications on January 26.
This research predicts a special molecular mechanism based on the docking simulation between PETase and a PET alternative mimic substrate. They constructed the variant for IsPETase with enhanced PET-degrading activity using structural-based protein engineering.
It is expected that the new approaches in this research could contribute to further study of other enzymes capable of degrading not only PET but other plastics, as well.
After use, PET causes environmental contamination issues due to its non-biodegradability. Conventionally, PET is disposed of in landfills, using incineration, and sometimes recycling using chemical methods, which induces additional environmental pollution. Therefore, highly-efficient PET degrading enzymes and bio-based, eco-friendly methods are desirable.
Recently, researchers isolated a new bacterial species, Ideonella sakaiensis, which can use PET as a carbon source. The PETase of I. sakaiensis (IsPETase) can degrade PET with relatively higher success than other PET-degrading enzymes. However, the detailed enzyme mechanism has not been elucidated, hindering further studies.
The research teams investigated how the substrate binds to the enzyme and which differences in enzyme structure result in significantly higher PET degrading activity compared with other cutinases and esterases. Based on the 3-D structure and related biochemical studies, they successfully determined the basis of the PET degrading activity of IsPETase and suggested other enzymes that can degrade PET with a new phylogenetic tree. The team proposed that 4 MHET moieties are the most properly matched substrates due to a structural cleft even with the 10-20-mers for PET. This is meaningful in that it is the first docking simulation between PETase and PET, not its monomer.
Furthermore, they succeeded in developing a new variant with much higher PET-degrading activity using a crystal structure of this variant to show that the changed structure better accommodates PET substrates than wild-type PETase, which will lead to superior enzyme development and platform construction for microbial plastic recycling.
Professor Lee said, "Environmental pollution from plastics remains one of the greatest challenges worldwide with the increasing consumption of plastics. We successfully constructed a new, superior PET-degrading variant with the determination of a crystal structure of PETase and its degrading molecular mechanism. This novel technology will help further studies to engineer more superior enzymes with high efficiency in degrading."
More information: Seongjoon Joo et al, Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation, Nature Communications (2018). DOI: 10.1038/s41467-018-02881-1
Journal information: Science , Nature Communications