Highly sensitive method to detect potential cancer biomarker
Exosomes regulate intercellular communication in cancers, and are drawing attention as a potential cancer biomarker. A Japanese research group has developed a highly sensitive method for detecting these exosomes that could potentially contribute to early detection of cancer.
The research group was focused around faculty in the Kobe University Medical Device Fabrication Engineering Center, led by Professor Toshifumi Takeuchi (Kobe University Graduate School of Engineering) and Professor Ryohei Sasaki (Kobe University Hospital), in collaboration with Kazuyuki Hamada (System Instruments Co., Ltd.). The findings were published on January 3 in Angewandte Chemie International Edition, and featured on the inner cover of the journal (Figure 1).
Exosomes are membrane vesicles consisting of lipid bilayer that measure only about 100 nanometers and are discharged from various cells. Exosomes secreted from cancer cells are closely linked to cancer migration, invasion, and metastasis. By examining these cancer-secreted exosomes, we may be able to develop a new method for early detection of cancer.
Current exosome analysis methods involve complicated and time-consuming procedures, including a combination of ultracentrifugation and affinity separation. If we could differentiate cancer-secreted exosomes from other exosomes in body fluids without complicated pretreatments, this would enable a simple test for early detection of cancer.
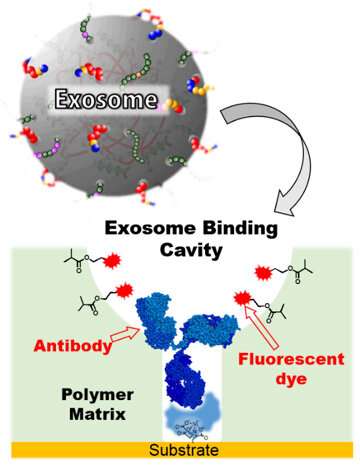
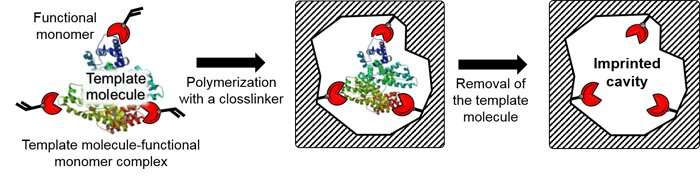
Using molecular imprinting technology (Figure 2), the team prepared an artificial polymer film of about 30 nanometers, possessing cavities the size of an exosome on top of the glass substrate (an exosome-binding cavity). Then, using their own post-imprinting technology, the researchers selectively introduced antibodies that can recognize membrane proteins on the surface of exosomes, and fluorescent molecules that can report exosome binding information. This resulted in an exosome-sensing chip (Figure 3), a groundbreaking, highly-sensitive fluorescence detection chip combining antibodies and artificial materials. It recognizes the membrane proteins on exosomes, captures the exosomes, and reports the information via changes in fluorescence.
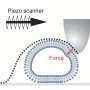
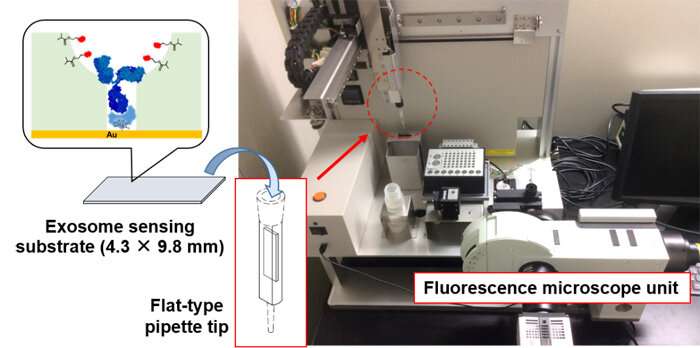
To simplify exosome analysis using this fluorescence-based sensing platform, the team collaborated with System Instruments Co., Ltd. to create a device that automatizes the whole analysis process (Figure 4). This instrument is a highly sensitive CMOS camera-mounted fluorescence microscope equipped with a 3-D robot arm. With a custom-made flat-type pipette tip inserted with the exosome sensing substrate, it can automatically carry out suction, discharge, fluorescence detection and substrate cleaning of the samples. Using this equipment, with no pretreatment and within 10 minutes, the team achieved unprecedented high-sensitivity detection of 6 picograms per milliliter. This technology enables us to detect about 150 exosomes in 10 microliters, a level of sensitivity that far surpasses previously reported measuring techniques.
For early detection of cancer, we need pre-checks before the disease takes hold, but currently the complexity of these checks means that only a small proportion of people are screened. The instrument developed in this study makes it easier to analyze the exosomes in bodily fluids. "Going forward, we will collect clinical samples on a large scale and carry out exosome analysis," comments Professor Takeuchi. "If it can be verified that analyzing cancer-secreted exosomes are helpful in cancer screenings, this will contribute to increasing the number of screenings. It can also be applied to cancer tests, treatment effectiveness, predictions of cancer transferal, and post-treatment prognoses."
More information: Kisho Mori et al. A Pretreatment-Free, Polymer-Based Platform Prepared by Molecular Imprinting and Post-Imprinting Modifications for Sensing Intact Exosomes, Angewandte Chemie International Edition (2018). DOI: 10.1002/anie.201811142
Journal information: Angewandte Chemie International Edition
Provided by Kobe University