How much are we learning? Natural selection is science's best critic

In 2003, the Human Genome Project revealed to the world the three billion chemical units within human DNA. Since that time, scientists have designed many ways to organize and assess this overwhelmingly large amount of information. Now, scientists at Cold Spring Harbor Laboratory (CSHL) have determined that evolution can help guide these efforts.
Researchers have already concluded that a mere one percent of the human genome is made up of the genes that make the proteins our bodies need to grow and function. However, they've also learned that roughly five percent of the human genome has remained the same, or been conserved, over countless generations of mutation and evolution.
"That suggests that an extra four percent of the genome is doing something that's really important, even though we don't know exactly what that is," explained Adam Siepel, a computational biologist and professor at CSHL.
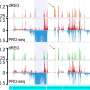
To solve the mystery of the four percent, scientists have spent more than a decade developing powerful methods to look for distinct functions among various bits of the genome. And, to understand what influences the genome has upon an organism, they've had to look to evidence from the epigenome. The epigenome is a universe of chemical compounds that attach themselves to DNA, influencing how and when parts of the genome are used by cells.
Searching for patterns among epigenomic factors has allowed scientists to guess where important parts of the genome may be and if they share biological function. However, this is no more certain than trying to determine the significance of a scene in a play by seeing only the props and costumes involved.
"This uncertainty about the true biological significance of many epigenomic measurements is a critical barrier not only for interpretation of the available data, but also for prospective decisions about how much new data to collect, of what type, and in what combinations," Siepel and his colleague Brad Gulko explained in the latest publication of Nature Genetics.
The Siepel lab has found a way around this barrier.
"So my lab and I decided to come at this from a different angle," added Siepel. "We asked, 'What if we let evolution do the work of telling us how much of the genome is important?' and, 'How much do we learn from each epigenomic data set?'"
The researchers used data from modern human populations to find evidence of recent natural selection. Then, they compared the genomes of humans and chimpanzees to get information that goes back five to seven million years to the divergence of humans from our great ape cousins.
"This allowed us to sort of chart how strong natural selection was during that whole period of time," Siepel explained.
The result was a way to guide future research. Siepel and his colleagues clustered sites within the genome based upon epigenomic features and how consequential each site has been for the survival of our species, according to evolutionary history. The resulting scores for each feature were then aggregated to create "fitness consequence maps," or FitCons maps.
If natural selection has been a powerful influence on a site in the genome—preserving it for countless generations despite mutation and evolution—this part of the genome should be important for survival. Moreover, if an epigenomic analysis identifies more of these conserved sites than not, then it will prove to be an informative study.
Siepel hopes that his fellow researchers will be able to reference FitCons to help determine which epigenetic markers or combinations of markers can prove the most informative for further investigation.
"This is an effort to try to see what we can learn by considering evolutionary information alongside what we already know," he said.
More information: Brad Gulko et al, An evolutionary framework for measuring epigenomic information and estimating cell-type-specific fitness consequences, Nature Genetics (2018). DOI: 10.1038/s41588-018-0300-z
Journal information: Nature Genetics
Provided by Cold Spring Harbor Laboratory