New megalibrary approach proves useful for the rapid discovery of new materials

Different eras of civilization are defined by the discovery of new materials, as new materials drive new capabilities. And yet, identifying the best material for a given application—catalysts, light-harvesting structures, biodiagnostic labels, pharmaceuticals and electronic devices—is traditionally a slow and daunting task. The options are nearly infinite, particularly at the nanoscale (a nanometer is one-billionth of a meter) where material properties—optical, structural, electrical, mechanical and chemical—can significantly change, even at a fixed composition.
A new study published this week in the Proceedings of the National Academy of Sciences (PNAS) supports the efficacy of a potentially revolutionary new tool developed at Northwestern University to rapidly test millions (even billions) of nanoparticles to determine the best for a specific use.
"When utilizing traditional methods to identify new materials, we have barely scratched the surface of what is possible," said Northwestern's Chad A. Mirkin, the study's corresponding author and a world leader in nanotechnology research and its applications. "This research provides proof-of-concept—that this powerful approach to discovery science works."
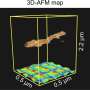
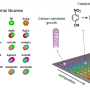
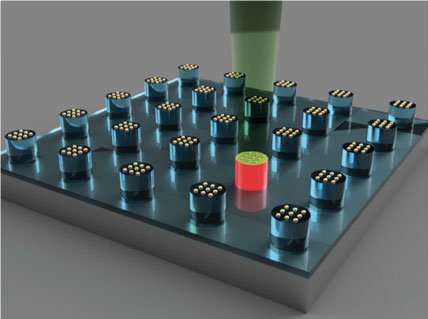
The novel tool utilizes a combinatorial library, or megalibrary, of nanoparticles in a very controlled way. (A combinatorial library is a collection of systematically varied structures encoded at specific sites on a surface). The libraries are created using Mirkin's Polymer Pen Lithography (PPL) technique, which relies on arrays (sets of data elements) with hundreds of thousands of pyramidal tips to deposit individual polymer "dots" of various sizes and composition, each loaded with different metal salts of interest, onto a surface. Once heated, these dots are reduced to metal atoms forming a single nanoparticle at fixed composition and size.
"By going small, we create two advantages in high throughput materials discovery," said Mirkin, the George B. Rathmann Professor of Chemistry in the Weinberg College of Arts and Sciences; professor of chemical and biological engineering, biomedical engineering and materials science and engineering in the McCormick School of Engineering; and executive director of Northwestern's International Institute for Nanotechnology (IIN). "First, we can pack millions of features into square-centimeter areas, creating a path for making the largest and most complex libraries, to date. Second, by working at the sub-100 nanometer-length scale, size can become a library parameter, and much of the action, for example, in the field of catalysis, is on this length scale."
The new study is a partnership between Northwestern's IIN and the Air Force Research Laboratory as part of the U.S. Air Force Center of Excellence for Advanced Bioprogrammable Nanomaterials at Northwestern. The team utilized a megalibrary and an in situ Raman spectroscopy-based screening technique called ARES to identify Au3Cu (a gold-copper composition) as a new catalyst for synthesizing single-walled carbon nanotubes. (ARES was developed by Benji Maruyama, leader, Flexible Materials and Processes Research Team, Materials & Manufacturing Directorate, Air Force Research Laboratory, and Rahul Rao, research scientist, Air Force Research Laboratory and UES, Inc.)
Carbon nanotubes are light, flexible and stronger-than-steel molecules used for energy storage, drug delivery and property-enhancing additives for many plastic materials. The screening process took less than one week to complete and is thousands of times faster than conventional screening methods.
"We were able to rapidly zero in on an optimal composition that produced the highest nanotube yield much faster than using conventional methods," said Maruyama, a study co-author. "The findings suggest we may have the ultimate discovery tool—a potential game changer in materials discovery."
More information: Edward J. Kluender et al, Catalyst discovery through megalibraries of nanomaterials, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1815358116
Journal information: Proceedings of the National Academy of Sciences
Provided by Northwestern University