Red-hued yeasts hold clues to producing better biofuels
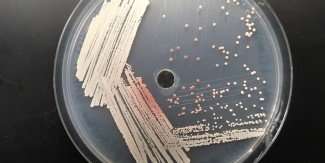
A compound that has scientists seeing red may hold the key to engineering yeasts that produce better biofuels.
A red pigment called pulcherrimin, naturally produced by several strains of wild yeasts, is synthesized in part through the same biochemical pathway that researchers hope to use to improve production of isobutanol, a promising biofuel alternative to ethanol. In research published this week (Oct. 8, 2018) in the Proceedings of the National Academy of Sciences, a team based at the University of Wisconsin–Madison and the Great Lakes Bioenergy Research Center describe the genetic machinery that yeasts use to make pulcherrimin, which binds iron, an essential nutrient. The work is a key step toward harnessing the synthesis pathway for large-scale production of isobutanol as a biofuel.
"Compared to first-generation biofuels, such as ethanol, isobutanol has a higher energy content, blends better with gasoline, causes less corrosion, and is more compatible with existing engine technology," says GLBRC researcher Chris Todd Hittinger, a UW–Madison genetics professor who led the research. "Nonetheless, considerable barriers remain to producing this fuel sustainably from dedicated energy crops."
Yeasts typically do not produce much isobutanol under normal conditions, says David Krause, a postdoctoral fellow with GLBRC and the lead author of the new study. Most commonly studied species produce ethanol during fermentation. But since the early steps of isobutanol synthesis are the same as those used to make pulcherrimin, yeasts that naturally produce the pigment – readily identifiable by their distinctive red hue – caught the researchers' eyes.
"Our thought is that these yeasts that are making pulcherrimin may be primed in a way to make more isobutanol," says Krause. "We want to use some of these yeast species that are already putting more carbon into these pathways and see if we can get them to turn that into isobutanol instead of pulcherrimin."
One challenge, though, was that not much was known about pulcherrimin, including how yeasts made it. The limited research available on the molecule focused on its chemical and antimicrobial properties. And the most common lab yeast species, Saccharomyces cerevisiae, does not make it at all.
The researchers used comparative genomics spanning 90 yeast species to identify the genes involved in pulcherrimin production. They found a cluster of four genes, which they named PUL1-4, that seem to play complementary roles. Through extensive genetic characterization, they determined that PUL1 and PUL2 are required to make the molecule, while PUL3 and PUL4 appear to help the yeast transport it and regulate its production.
The discovery was surprising in part because it marks the first report of a gene cluster in budding yeasts responsible for producing a type of compound known as a secondary metabolite. Many secondary metabolites have valuable functions as antibiotics, toxins or signaling molecules. While many such molecules are produced by filamentous fungi and bacteria, the new research suggests some budding yeasts make secondary metabolites as well.
"Studying diverse genomes can lead to discoveries and new biological insights... We were able to learn more about genes in S. cerevisiae through the lenses of some of these lesser-known species," said Chris Hittinger.
Another surprising aspect of the study was the finding that many yeast species that do not make pulcherrimin – including S. cerevisiae – nonetheless have working PUL3 and PUL4 genes. Patterns across many yeast lineages suggest that retaining these genes allows some species to capitalize on the pulcherrimin made by others, Krause explains.
"There can be an evolutionary trend toward organisms that dispense with the ability to produce a molecule but still are able to use it," he says. "So, their neighbors are making pulcherrimin, and they're able to use it without having to incur the costs of making it."
The findings also highlight the value of stepping beyond traditional lab models.
"This work really shows how studying diverse genomes can lead to discoveries and new biological insights," says Hittinger. "Focusing on a single organism can give us an incomplete picture of a complex biological process. At the same time, we were able to learn more about genes in S. cerevisiae through the lenses of some of these lesser-known species."
With a better understanding of the steps involved in pulcherrimin production, the researchers are now poised to try to tweak the production machinery and turn it to making isobutanol instead. "This research is a starting point for taking what we've learned about pulcherrimin and applying it to biofuels," Krause says.
More information: David J. Krause et al. Functional and evolutionary characterization of a secondary metabolite gene cluster in budding yeasts, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1806268115
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Wisconsin-Madison




















