Molecular virologist fights influenza at the molecular level
Molecular virologist Chad Petit, Ph.D., uses basic science to fight influenza—through experiments at the atomic level.
This includes a deadly poultry influenza virus in China called the H7N9 avian flu virus. Since 2013, H7N9 has infected 1,625 people, killing 623. While not highly contagious for humans, just three mutations could change that, turning H7N9 into the feared Disease X, the term health experts use for the next unknown cause of a worldwide epidemic.
In research to improve influenza therapies against H7N9 and other influenza strains, Petit and his University of Alabama at Birmingham colleagues have detailed the binding site and mechanism of inhibition for two small-molecule experimental inhibitors of influenza viruses. Their report is published in the Journal of Biological Chemistry.
The two experimental inhibitors studied by Petit, a UAB assistant professor of biochemistry and molecular genetics, are small molecules whose precise mechanism of action was unknown. The inhibitors target the function of a key influenza protein called NS1, which has multiple roles to block the body's immune response during influenza infection. Thus, NS1 is essential to the survival and adaptability of the influenza virus.
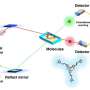
Petit and colleagues used nuclear magnetic resonance, or NMR, spectroscopy to probe interactions of the inhibitors with NS1. They first showed that the inhibitors—called A9 and A22—interacted with just one of the two independently folded domains of NS1, the NS1 effector domain.
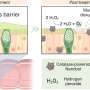
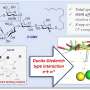
The researchers noted that the structures of both small-molecule inhibitors were very similar to a fragment of a host protein called CPSF30 that the NS1 effector domain binds in order to short-circuit the body's immune response. Therefore, the researchers hypothesized that A9 and A22 block influenza viral replication and block NS1 function by interfering with the interaction between the NS1 effector domain and CPSF30.
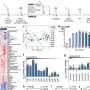
NMR data revealed the particular amino acids of the NS1 effector domain that are involved in inhibitor binding. The researchers—using two significantly different NS1 proteins from distinct influenza strains, including the H7N9 strain—showed that similar sequences of amino acids in the two NS1 proteins were involved in inhibitor binding.
The 1918 "Spanish" flu NS1 protein
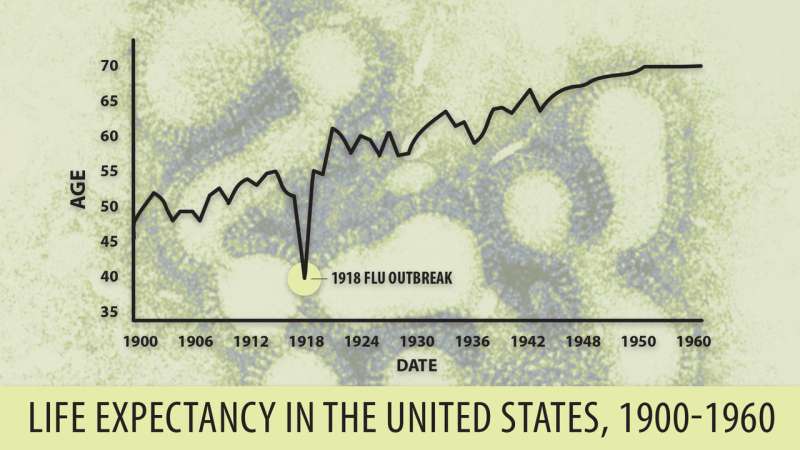
Besides the Chinese H7N9 NS1, the other NS1 protein tested was the NS1 effector domain from the 1918 "Spanish" flu, which infected one-third of the world's population a century ago and killed 50 million to 100 million people.
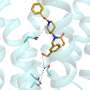
The UAB researchers then used X-ray crystallography, led by UAB Microbiology assistant professor Todd Green, Ph.D., to determine the three-dimensional structure of the NS1 effector domain from the 1918 "Spanish" flu. This allowed them to map the A9/A22-binding site onto that structure, which confirmed their hypothesis—A9 and A22 interact with the NS1 effector domain hydrophobic pocket that is known to bind the host protein CPSF30.
The crystallography data also showed that the NS1 effector domain is able to dimerize, using an interface different from two other known dimers of the NS1 effector domain. Biological significance of this new dimer form is unknown.
"Altogether, our findings provide strong evidence for the mechanism of action of two anti-influenza compounds that target NS1, and the findings contribute significant structural insights into NS1 that we hope will promote and inform the development and optimization of influenza therapies based on A9 and A22," Petit said.
The need for novel antiviral compounds is great. Each year, influenza strains kill 250,000 to 500,000 people worldwide, and the virus is noted for quick changes to produce pandemic strains that few people have immunity against. Viral resistance has limited the effectiveness of several earlier antiviral compounds that were developed to treat influenza.
More information: Alex B. Kleinpeter et al, Structural analyses reveal the mechanism of inhibition of influenza virus NS1 by two antiviral compounds, Journal of Biological Chemistry (2018). DOI: 10.1074/jbc.RA118.004012
Journal information: Journal of Biological Chemistry
Provided by University of Alabama at Birmingham