Screening technique detects special messenger RNA molecules that sense and respond to environmental signals
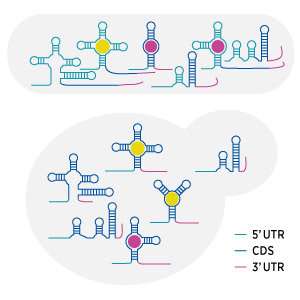
A*STAR researchers have developed a rapid, high-throughput screening procedure that can detect RNA aptamers—special messenger RNA molecules (mRNAs) able to sense and respond to particular cellular chemicals—in a sea of normal mRNA extracted from cells.
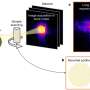
The process, developed by a team led by Yue Wan of the A*STAR Genome Institute of Singapore, is known as PARCEL or 'Parallel Analysis of RNA Conformations Exposed to Ligand binding'. Wan hopes the technique will become a standard tool in gene regulation research.
"PARCEL provides a very rapid way to find RNA aptamers in almost any cell type," she says.
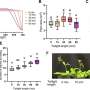
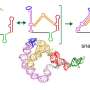
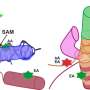
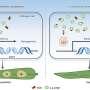
RNA aptamers are very specific: they sense just one chemical by binding to it, which can alter their shape. By comparing the shapes of mRNA molecules extracted from cells, grown in either the presence or absence of the target chemical, PARCEL can spot the few mRNAs able to sense that chemical.
The chemicals sensed are usually metabolites that play a significant role in the cell's biochemistry, says Wan.
To date, PARCEL has found 58 new RNA aptamers—a significant addition to the few natural RNA aptamers that were already known.
Importantly, a few mRNA aptamers have been previously shown not only to sense a metabolite but also to initiate a response. These are called riboswitches. "The riboswitch tells the cell 'Hey I've sensed a molecule—so now take this action'", says Wan.
Messenger RNA molecules normally carry a coded 'message' from a gene to the cell's protein factory, requesting production of a particular protein. So, Wan notes, one possible action for the cell is to alter the production rate of that protein.
Before PARCEL, riboswitches had been found in bacteria using a computational approach. But few were known to exist in eukaryotes, single-celled micro-organisms with a nucleus. Wan and her team hypothesized that eukaryotes might have riboswitches sensitive to a key metabolite—vitamin B2.
"Vitamin B2 is a very important metabolite and in bacteria there are many riboswitches that respond to it. We decided to use PARCEL to screen for them in eukaryotes."
PARCEL revealed two new riboswitches in eukaryotes—specifically, yeast cells—that either enhanced or reduced protein production depending on the vitamin B2 level, plus another bacterial vitamin B2 riboswitch.
Wan anticipates using PARCEL will lead to many more such discoveries.
More information: Sidika Tapsin et al. Genome-wide identification of natural RNA aptamers in prokaryotes and eukaryotes, Nature Communications (2018). DOI: 10.1038/s41467-018-03675-1
Journal information: Nature Communications