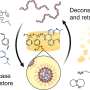
How molecular riboswitches work in bacteria
Many bacteria have molecular control elements via which they can switch genes on and off. These riboswitches also open up new options in the development of antibiotics or the detection and decomposition of environmental toxins. Researchers of Karlsruhe Institute of Technology (KIT), Heidelberg University, and Freie Universität Berlin have now used light optical microscopy of single molecules to fundamentally study the way riboswitches work. This is reported in Nature Chemical Biology.
Riboswitches are located on the messenger ribonucleic acid (mRNA) that transports genetic information to the site of protein biosynthesis. A riboswitch consists of a sensor measuring the concentration of a small metabolic molecule and an effector controlling gene expression and, hence, synthesis of a protein. As riboswitches exist in many bacterial pathogens, they represent important targets in the development of new antibiotics. Other applications are possible in synthetic biology. For example, bacteria can be genetically modified with riboswitches to detect and decompose low-molecular environmental toxins, such as herbicides. However, fundamental understanding of the processes underlying the function of riboswitches is required. The work presented in Nature Chemical Biology is an essential contribution in this respect.
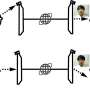
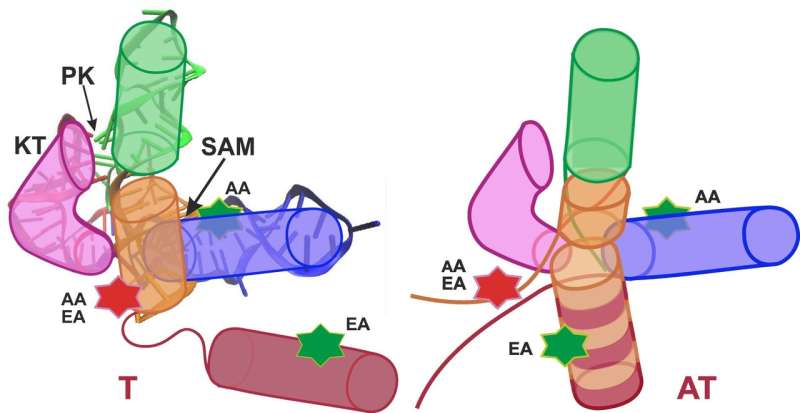
The research groups of Professor Gerd Ulrich Nienhaus of KIT and Professor Andres Jäschke of Heidelberg University studied the S-adenosyl-L-methionine (SAM)-I riboswitch. "Attachment of the SAM molecule to this riboswitch causes the conformation, that is the spatial arrangement of atoms, to change from the anti-terminator (AT) to the terminator (T) structure," Nienhaus explains. "As a result, gene expression is switched off."
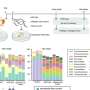
First, the scientists in Heidelberg synthesized SAM-I riboswitches and specifically marked them with two fluorescent dyes each at different points. The researchers of KIT then studied these RNA molecules at high spatial and temporal resolution using highly sensitive light microscopes measuring fluorescent emission of single-dye molecules. By means of Förster resonance energy transfer (FRET) experiments, conformation dynamics were determined directly. For this purpose, laser radiation is used to make a green dye emit light. If a red dye is located in the vicinity, it can take over the excitation energy of the green dye and emit light itself.
The probability of energy transfer strongly depends on the distance of the dyes from each other. Structural changes of a molecule to which the dyes are specifically attached can be observed directly via emission of the red dye. Light emission is extremely weak, requiring complex data analysis methods based on hidden Markov modeling. Professor Bettina Keller of the Institute of Chemistry and Biochemistry of Freie Universität Berlin developed the methods especially for this type of experiment in order to distinguish time-dependent light emission signals from noise.
In their analysis, the researchers distinguished two conformations (T and AT) of the SAM-I riboswitch, and a total of four conformations (T1, T2, AT1, and AT2). Surprisingly, the riboswitch did not fully switch between the T and AT structures in the presence and absence of SAM, as had been expected, but fluctuated permanently between all states—only weightings were shifted. A result important to the biological function was that structure fluctuations observed with an attached SAM were far quicker than without SAM. As the riboswitch sequence on the messenger RNA is located directly in front of the gene to be controlled, the RNA molecule has to form a T structure (switch off) as quickly as possible after synthesis in the presence of SAM in order to prevent subsequent transcription of the gene to be controlled. Acceleration of structure fluctuations by SAM attachment thus ensures sufficiently quick formation of a T structure. "Consequently, dynamics of the SAM-I riboswitch play an important role for its function," Nienhaus says. "These detailed insights into the functioning of a biomolecule are the results of an interdisciplinary approach of physics, biotechnology, and theoretical chemistry."
More information: Christoph Manz et al. Single-molecule FRET reveals the energy landscape of the full-length SAM-I riboswitch, Nature Chemical Biology (2017). DOI: 10.1038/nchembio.2476
Journal information: Nature Chemical Biology
Provided by Karlsruhe Institute of Technology