Combined X-ray and fluorescence microscope reveals unseen molecular details
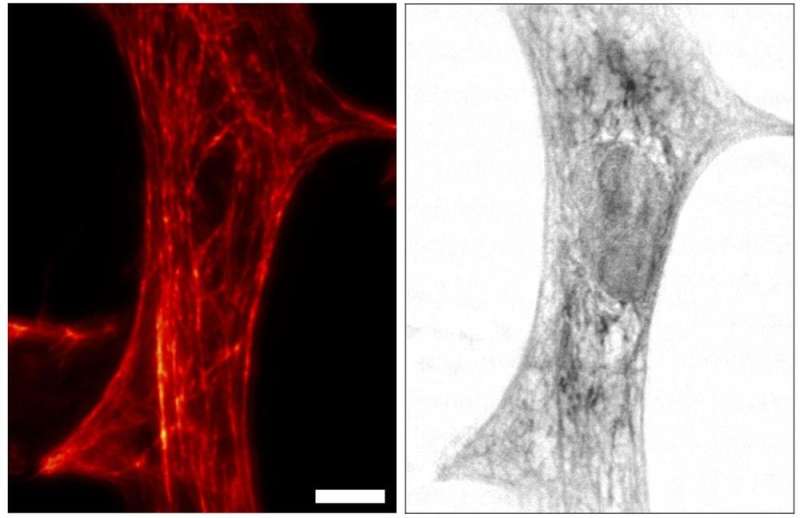
A research team from the University of Göttingen has commissioned a worldwide unique microscope combination at DESY's X-ray source PETRA III to gain novel insights into biological cells. The team led by Tim Salditt and Sarah Köster describes the combined X-ray and optical fluorescence microscope in the journal Nature Communications. To test the performance of the device installed at DESY's measuring station P10, the scientists investigated heart muscle cells with their new method.
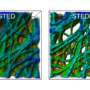
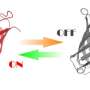
Modern light microscopy provides with ever sharper images important new insights into the interior processes of biological cells, but highest resolution is obtained only for the fraction of biomolecules which emit fluorescence light. For this purpose, small fluorescent markers have to be first attached to the molecules of interest, for example proteins or DNA. The controlled switching of the fluorescent dye in the so-called STED (stimulated emission depletion) microscope then enables highest resolution down to a few billionth of a meter, according to principle of optical switching between on- and off-state introduced by Nobel prize winner Stefan Hell from Göttingen.
"But how can we get sharp images of all cellular components, including those molecules to which fluorescent markers cannot be attached," asks Salditt. "How can we illuminate the 'dark background' of all unlabelled molecules, in which the specifically labelled fluorescent biomolecules are embedded?"
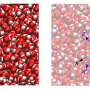
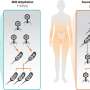
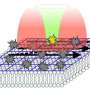
Salditt's and Köster's team has now combined a STED- and an X-ray microscope, which can quasi simultaneously map fluorescence and the density distribution of the total of cellular components in the cell. "In addition, X-ray diffraction experiments, which are well known from crystallography, can also be carried out at precisely controlled positions in the cell," explains co-author Michael Sprung, head of the measuring station P10 where the new device has been installed.
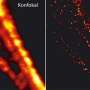
"With this novel X-ray/STED microscope we have visualised a network of protein filaments in heart muscle cells in STED mode first. The cells were then also imaged by X-ray holography to cover the spatial distribution of mass density in the entire cell, including all of its components," explains Marten Bernhardt, lead author of the article. "By using complementary contrast we aim at a more complete understanding of the structure underlying contractibility and force generation in the cells," adds Salditt. "In future, we want to apply this also to observe dynamic processes in living cells," explains Köster, spokesperson of the collaborative research centre Collective behaviour of soft and biologcal matter of the German Science Foundation (DFG), which provides the research framework of the experiments.
More information: M. Bernhardt et al. Correlative microscopy approach for biology using X-ray holography, X-ray scanning diffraction and STED microscopy, Nature Communications (2018). DOI: 10.1038/s41467-018-05885-z
Journal information: Nature Communications
Provided by Helmholtz Association of German Research Centres