New genetics tool helps answer evolutionary questions

The age of big data is here. Thanks to innovations in genetic sequencing technology, scientists can now generate massive datasets describing the genomes of Earth's diverse set of species. This ever-growing genomic encyclopedia has the capacity to reveal the forces shaping complex patterns of genetic variation between individuals, populations and species—if scientists can only unlock its secrets.
Developing cutting-edge statistical tools that can handle these massive new datasets is a piece of the research puzzle, and new research from Michigan State University has just added a new tool for the modern genomic toolbox.
The method, called "conStruct" and featured in the current issue of Genetics, allows researchers to analyze complex patterns of genetic variation in large datasets with broad geographic sampling. It overcomes major shortcomings of previous methods and is free and publicly available worldwide.
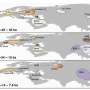
"One of the first steps in the analysis of these genomic datasets is to describe and categorize variation into discrete populations, like you might find in range maps in a field guide," said Gideon Bradburd, MSU population geneticist and lead author. "What often determines relatedness is geography. If you sample two organisms separated by a large distance, you often have to go farther back into the history of their pedigrees to find a shared ancestor."
This leads to isolation by distance, a pattern that creates statistical challenges for anyone interested in cleanly describing variation within and between groups in their own study system, he added.
In the paper, Bradburd and his colleagues illustrate the utility of their new approach by applying it to genomic data collected on North American bears and poplar trees. For a better understanding, let's look at the poplars, which are distributed throughout the northern hemisphere. Different species of poplar can be found near each other, and, where they overlap, they frequently hybridize.
Using conStruct, the research team was able to review the degree to which hybridization between the two poplar species has happened. They were also able to determine whether the only significant population boundary fell along the species boundary, and if there was substructuring within the species.
"Understanding the genetic relatedness of individuals is central to many important research fields, including conservation biology, human medicine, evolution and ecology, and agriculture," Bradburd said. "With conStruct, scientists can home in on commonalities and discrepancies among populations with more accuracy. This can prove invaluable, especially in conservation efforts."
And, of course, these genomic patterns will offer additional insights into human evolution.
For the next phase of this research, Bradburd's team will attempt to take conStruct into the fourth dimension. They hope to add the ability to model historical or ancient DNA samples to learn how and why populations change—or are replaced by their neighbors—through time.
More information: Gideon S. Bradburd et al. Inferring Continuous and Discrete Population Genetic Structure Across Space, Genetics (2018). DOI: 10.1534/genetics.118.301333
Journal information: Genetics
Provided by Michigan State University