Searching the sea, and bacterial battles, for new antibiotics

Alexander Fleming's discovery of penicillin—the world's first natural antibiotic—is famously told as a story of serendipity: a petri dish growing bacteria was contaminated by mold, which secreted a substance to keep bacteria at bay. The lesson learned was that science can take advantage of chance encounters to change the world.
But perhaps the scientists that followed in Fleming's footsteps should have paid closer attention to a core aspect of his discovery: that it hinged on the relationship between microscopic competitors fighting for space.
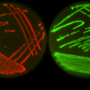
University of Wisconsin–Madison researchers are collaborating across colleges and departments to relearn the lessons of penicillin. They are stepping beyond an era when microbes were grown alone in search of new antibiotics by growing different species together. Their goal is to stimulate natural defenses against old foes, like what took place when the penicillium mold attacked staphylococcus bacteria in Fleming's lab cultures. These coculturing techniques aim to recreate aspects of real ecosystems to spur dormant and hidden antibiotic capacities into action.
After years improving this relatively new method, scientists at the UW–Madison School of Pharmacy and the College of Agricultural and Life Sciences discovered the new antibiotic keyicin, a demonstration of the technique's effectiveness. Researchers say this discovery would not have been possible without a cross-college collaboration going back nearly a decade.
The microbes that give us most of our antibiotics never grow alone in nature. Yet they have that luxury in the lab, where scientists grow bacteria or fungi in isolation to study them one at a time. Many life-saving antibiotics were discovered under these conditions. But over time those discoveries waned, while pathogens began developing resistance to existing drugs.
"The well had run dry," says Tim Bugni, a chemist by training and a professor of pharmacy at UW–Madison who was the senior author of the paper announcing keyicin, published in 2017. "In the '90s, most pharmaceutical companies abandoned this area of research. Starting in 2000, genomics really started taking off."
The genomic era revealed a tantalizing opportunity: DNA sequencing showed that many bacteria had troves of genes for making novel antibiotics. They were just never activated. Even the most creative laboratory conditions could not induce the microbes to tap into this arsenal of new chemicals.
When Bugni arrived at UW–Madison in 2009, he soon began working with Cameron Currie, a professor of bacteriology. The two shared an interest in partnerships between microbes and animals and in antibiotic research.
"A lot of these silent genetic capacities for producing antimicrobial compounds are linked to the ecological role they play," says Currie, a co-author of the keyicin paper. Professor of Pharmacy Lingjun Li also contributed to the work, which was led by Bugni's former graduate student Navid Adnani. Collaborators at the University of Minnesota, Yumanity Therapeutics and Bruker Daltonics also contributed.
"Given that producing antibiotics is energetically costly to bacteria, if they are using them in an ecological framework, to inhibit a pathogen or competitor, it makes sense from an evolutionary perspective to only do so when they get a signal from the target organism, instead of continually cranking it out," says Currie.
In theory, a competing microbe provides that missing signal. In response to the threat, bacteria turn on their once-silent genes, pumping out a previously uncharacterized antibiotic. The researchers discovered keyicin when the bacterium Micromonospora was challenged with Rhodococcus. Over time, the keyicin produced by Micromonospora helped it take over the culture.
Both strains of bacteria came from the ocean, where they are associated with invertebrates. A sizable portion of existing antibiotics were discovered in bacteria that live in the soil. But continued work with these terrestrial bacteria has discovered the same drugs over and over again. Bugni, who specializes in marine microbes, says tapping into this relatively uncharted ecosystem gives scientists a better chance of avoiding this "rediscovery problem," which plagues antibiotic research.
"There's a lot of unexplored bacterial diversity in the marine environment," Bugni says.
The coculturing work is funded by a Center of Excellence for Translational Research grant from the National Institutes of Health. David Andes, a professor and chief of infectious diseases at the University of Wisconsin School of Medicine and Public Health, leads the grant, which Currie and Bugni are members of. While Bugni focuses on marine bacteria, Currie specializes in microbes associated with insects on land.
The team is assessing keyicin for its therapeutic potential in animals. (Most new antibiotics face considerable hurdles to being used in humans, but only more research will tell.) In the meantime, researchers say the proof of concept provided by the discovery of keyicin suggests that coculturing will continue to provide more new antibiotic candidates.
This approach requires evolutionary, biological, chemical and medical expertise directed at an increasingly complex problem.
"This kind of interdisciplinary work is absolutely critical to be successful in this realm," says Currie.
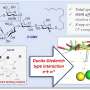
More information: Navid Adnani et al. Coculture of Marine Invertebrate-Associated Bacteria and Interdisciplinary Technologies Enable Biosynthesis and Discovery of a New Antibiotic, Keyicin, ACS Chemical Biology (2017). DOI: 10.1021/acschembio.7b00688
Journal information: ACS Chemical Biology
Provided by University of Wisconsin-Madison