Stone Age hepatitis B virus decoded

An international team of scientists led by researchers at the Max Planck Institute for the Science of Human History and the University of Kiel has successfully reconstructed genomes from Stone Age and Medieval European strains of the hepatitis B virus. This unprecedented recovery of ancient virus DNA indicates that hepatitis B was circulating in Europe at least 7000 years ago. While the ancient virus is similar to its modern counterparts, the strains represent a distinct lineage that has likely gone extinct and is most closely related to chimpanzee and gorilla viruses.
The hepatitis B virus (HBV) is one of the most widespread human pathogens known today, affecting over 250 million people worldwide. However, its origin and evolutionary history remain unclear. Studying the evolution and history of the virus has to date been especially difficult, because until now viral DNA had not been successfully recovered from prehistoric samples. In the present study, which has been accepted for publication in the journal eLife and is due to be published on May 10, 2018, an international team of researchers led by the Max Planck Institute for the Science of Human History and the Institute of Clinical Molecular Biology at Kiel University, not only recovered ancient viral DNA from skeletons but also reconstructed the genomes of three strains of HBV.
The ancient history of hepatitis B
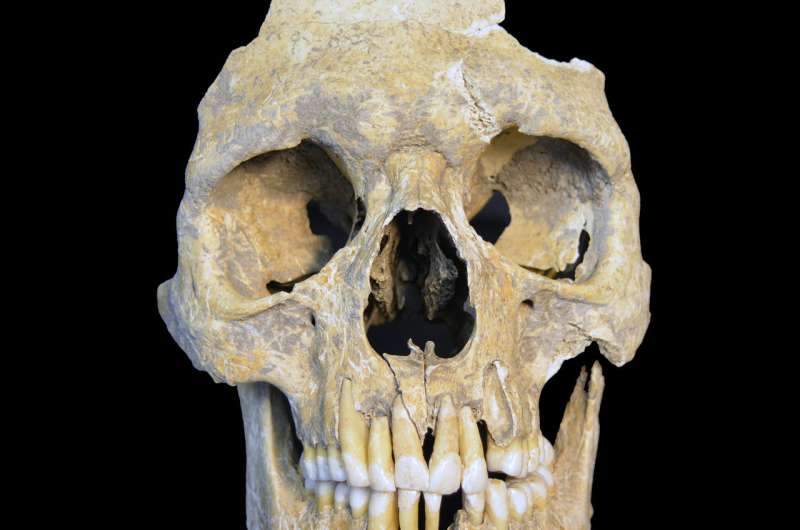
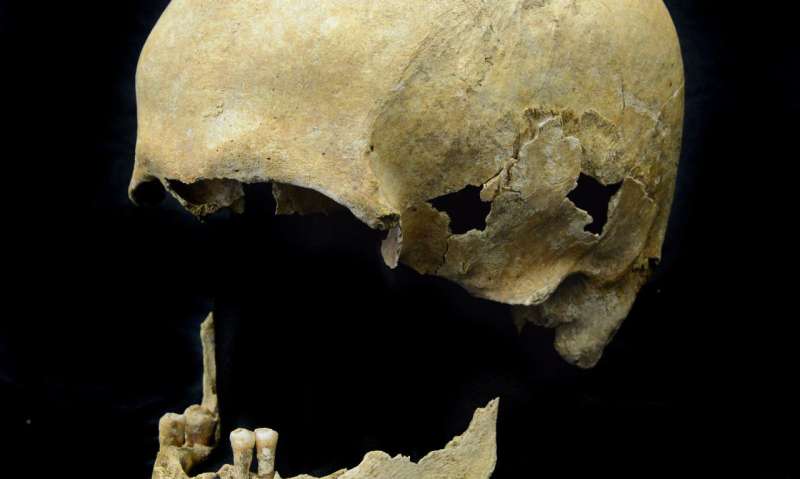
For this study, the researchers analyzed samples from the teeth of 53 skeletons excavated from Neolithic and medieval sites in Germany. The remains dated from around 5000 BC to 1200 AD. The researchers screened all samples for viral pathogens and detected ancient HBV in three of the individuals. Full HBV genomes were recovered from these samples, two of which were from the Neolithic period, dating to about 7000 and 5000 years ago, and one from the medieval period. The Neolithic genomes represent the by far oldest virus genomes reconstructed to date.
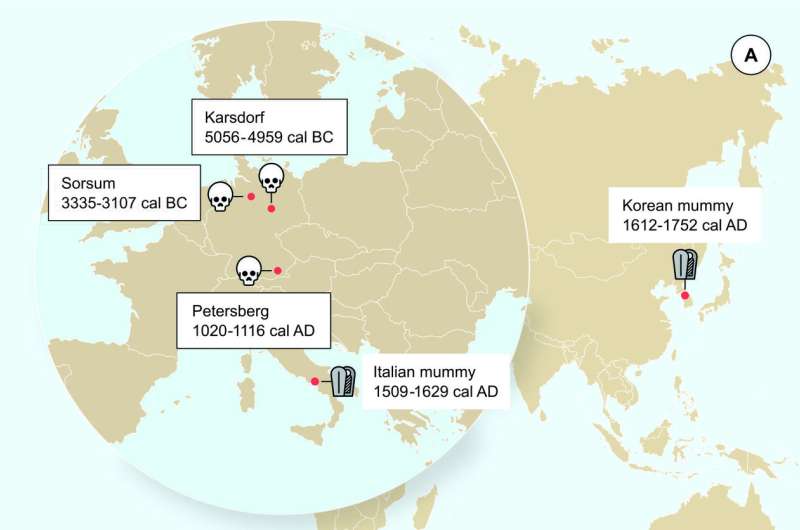
Interestingly, the ancient virus genomes appear to represent distinct lineages that have no close relatives today and possibly went extinct. The two Neolithic genomes, although recovered from individuals that lived 2000 years apart, were relatively similar to each other in comparison with modern strains, and were in fact more closely related to modern strains of HBV found in Chimpanzees and Gorillas. In contrast, the medieval HBV genome is more similar to modern strains, but still represents a separate lineage. This is the case even when it is compared to two previously published HBV genomes recovered from mummies dating to the 16th century. The HBV strains found in these mummies are closely related to modern strains, suggesting a surprising lack of change in the virus over the last 500 years. These findings point to a complicated history for the virus, which may have involved multiple cross-species transmission events.
Long and complicated evolution of one of today's most common viruses
"Taken together, our results demonstrate that HBV already existed in Europeans 7000 years ago and that its genomic structure closely resembled that of modern hepatitis B viruses, despite the differences observed," explains first author Ben Krause-Kyora, of the Max Planck Institute for the Science of Human History and Kiel University. "More ancient precursors, intermediates and modern strains of both human and non-human primate HBV strains need to be sequenced to disentangle the complex evolution of this virus," he adds.
Johannes Krause, senior author and director of the Department of Archaeogenetics at the Max Planck Institute for the Science of Human History, emphasizes the most important implication of the study. "Our results reveal the great potential of ancient DNA from human skeletons to allow us to study the evolution of blood-borne viruses. Previously, there was doubt as to whether we would ever be able to study these diseases directly in the past," he explains. "We now have a powerful tool to explore the deep evolutionary history of viral diseases."
More information: eLife, DOI: 10.7554/eLife.36666
Journal information: eLife
Provided by Max Planck Society