A molecular dance of phospholipid synthesis
The most abundant molecule in cell membranes is the lipid phosphatidylcholine (PC, commonly known as lecithin); accordingly, the enzymes responsible for synthesizing it are essential. Research published in the May 4 issue of the Journal of Biological Chemistry used computer simulations to gain insights into how one of these enzymes activates and shuts off PC production. These results could help researchers understand why small changes in this enzyme can lead to conditions like blindness and dwarfism.
Rosemary Cornell, a professor of molecular biology and biochemistry at Simon Fraser University in Canada, studies the enzyme CTP:phosphocholine cytidylyltransferase, or CCT. CCT sets the rate of PC production in cells by binding to cell membranes with low PC content. When bound to membranes, the CCT enzyme changes shape in a way that allows it to carry out the key rate-limiting step in PC synthesis. When the amount of PC making up the membrane increases, the CCT falls off the membrane, and PC production ceases.
"The membrane is this big macromolecular array with lots of different molecules in it," Cornell said. "How does this enzyme recognize that 'Oh, I should slow down because the PC content of the membrane is getting too high?'"
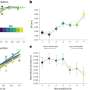
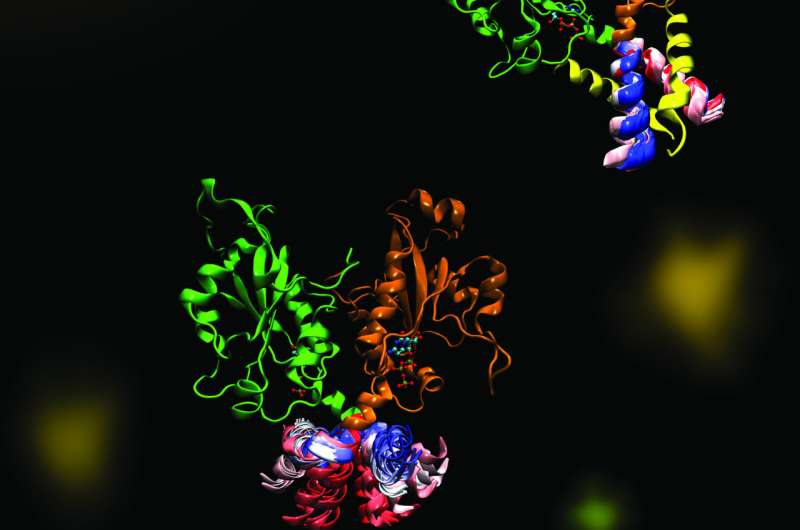
Cornell and her project team - a collaboration with Peter Tieleman and graduate student, Mohsen Ramezanpour at the University of Calgary and Jaeyong Lee and Svetla Taneva, research associates at SFU - thought that the answer must have to do with the dynamic changes in shape that the enzyme undergoes when it binds to a membrane. But these changes are difficult to capture with traditional structural biology methods like x-ray crystallography, which take a static snapshot of molecules. Instead, the team used computational simulations of molecular dynamics, which use information about the forces between every individual atom in a molecule to calculate the trajectories of the enzyme's moving parts.
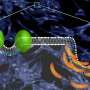
"What it looks like (when you visualize the output) is your big molecule dancing in front of your eyes," Cornell said. "We set up the molecular dynamics simulation not once, not twice, but 40 different (times). It took months and months just to do the computational parts and even more months trying to analyze the data afterward. We actually spent a lot of time once we got the data just looking on the screen at these dancing molecules."
The simulated dance of the CCT molecule showed that when the M-domain, the section of the enzyme that typically binds to the membrane, detaches from a membrane, it snags the active site of the enzyme, preventing it from carrying out its reaction. When the snagging segment was removed from the simulation, the team saw a dramatic bending motion in the docking site for the snagging element, and speculated that this bending would create a better enzyme active site for catalyzing the reaction when attached to a membrane. The team confirmed these mechanisms using biochemical laboratory experiments.
Interestingly, previous genetic studies had shown that mutations in the gene encoding CCT are responsible for rare conditions like spondylometaphyseal dysplasia with cone-rod dystrophy, which causes severe impairments in bone growth and vision, but it was unknown how these changes in the enzyme could lead to such dramatic consequences. Cornell hopes that understanding how the enzyme works could help researchers find out.
"If you have just one small change in CCT, then how is that going to make this whole process of synthesizing PC defective?" Cornell asks. "That's what we're studying right now."
More information: Mohsen Ramezanpour et al, An auto-inhibitory helix in CTP:phosphocholine cytidylyltransferase hijacks the catalytic residue and constrains a pliable, domain-bridging helix pair, Journal of Biological Chemistry (2018). DOI: 10.1074/jbc.RA118.002053
Journal information: Journal of Biological Chemistry