Bacterial 'gene swapping' sparks disease outbreaks
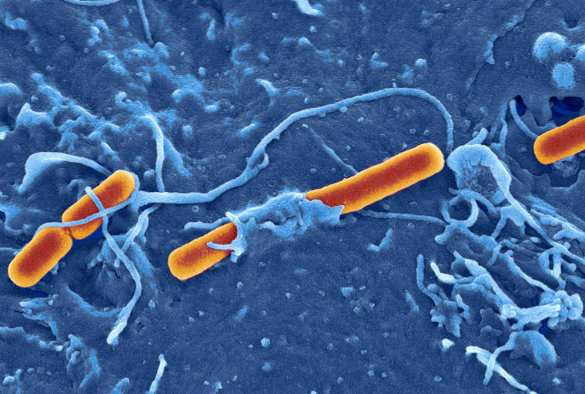
A new study by scientists at the University of Liverpool documents, for the first time, how the ability of bacteria to swap genetic material with each other can directly affect the emergence and spread of globally important infectious diseases.
Known as 'horizontal gene transfer', this phenomenon is understood to have played a role in developing the global antimicrobial resistance (AMR) crisis. However, the dynamics of AMR transfer through bacterial populations and its direct impact on human disease is poorly understood.
To learn more about this relationship, a team of researchers from the University of Liverpool, Wellcome Trust Sanger Institute and Public Health England, combined epidemiological information from cases together with whole genome sequencing analysis of bacteria from those cases.
The study, which is published in Nature Communications, looked at three epidemics in England of the globally important diarrhoeal pathogen Shigella between 2008 and 2014.
The findings found that English epidemics of these typically-rare pathogens were associated with resistance to the antibiotic azithromycin, and that many of the co-circulating Shigella strains carried the azithromycin resistance genes on the same plasmid.
Plasmids are small circular DNA molecules that can be transferred horizontally between bacteria. They contain the bacterium's genetic material but are separate from the cell's chromosomal DNA.
By combining this genomic information from the different Shigella strains with the epidemiological information about the outbreaks, the researchers were able to demonstrate that the transfer of the plasmid was facilitating new epidemics.
Dr. Kate Baker, from the University's Institute of Integrative Biology, said: "Through this study we've been able to show that horizontal gene transfer can rapidly facilitate new epidemics of important pathogens.
"This means that in all areas of AMR research, public health management and surveillance we need to be analysing our pathogen genomes in great detail to understand the epidemiology of antimicrobial resistance."
Antimicrobial resistance (AMR) is a global epidemic. In 2014 the World Health Organisation advised that the antibiotics we are reliant on are in danger of becoming obsolete. It is estimated that by the year 2050, deaths attributed to antimicrobial resistance will claim up to ten million lives per year, surpassing those lost to cancer.
More information: Kate S. Baker et al. Horizontal antimicrobial resistance transfer drives epidemics of multiple Shigella species, Nature Communications (2018). DOI: 10.1038/s41467-018-03949-8
Journal information: Nature Communications
Provided by University of Liverpool