A highly sensitive and multi-analytical system for hereditary kidney disease
Alport syndrome (AS) is a hereditary kidney disease caused by a genetic mutation leading to type IV collagen (Col4) abnormalities. Unfortunately, treatment through the correction of Col4 functionality has not yet been developed. Now, researchers from Kumamoto University in Japan have established a highly sensitive technology to assess Col4 functionality, paving the way to therapeutic drugs. This detection system reduces labor and time costs compared to conventional methods, and monitors the functionality of Col4 with higher sensitivity than ever before. The system also makes it possible to analyze several drug candidate compounds simultaneously.
ACE inhibitors and other medications for treating hypertension have been found to slow the progression of AS. However, this treatment only alleviates symptoms and cannot prevent the transition into end stage renal failure. It is therefore considered that fundamental treatment should focus on the cause of disease onset, an approach that is completely different from prior methods. The therapeutic strategy, in this case, would focus on normalizing the function of the causative protein with a candidate drug compound. If successful, this technique would greatly contribute to the treatment of other hereditary diseases.
To normalize the function of the causative protein with a candidate drug compound, it is necessary to efficiently screen and identify compounds that restore the function compromised by the genetic mutation. However, an analysis system to assess the functionality of causative proteins has not yet been developed. Such a system could be used to screen for therapeutic drug candidates. Thus, Kumamoto University researchers aimed to establish a new evaluation system (a compound screening system) just for that purpose.
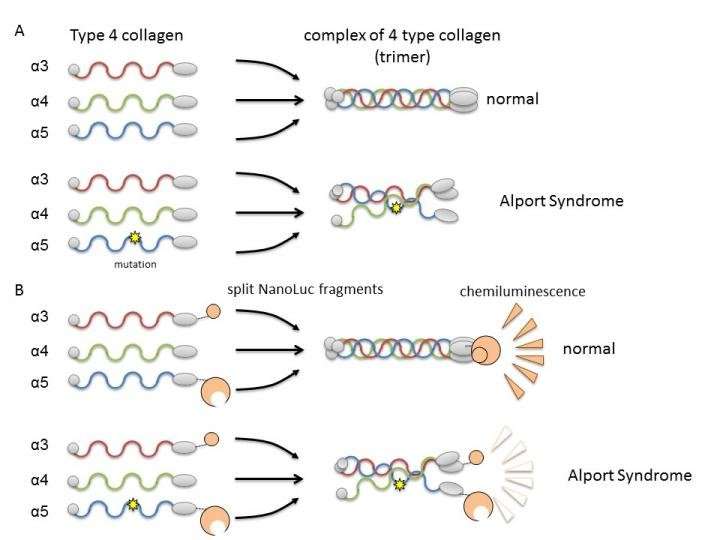
In Col4, three polypeptide chains (rod-like proteins), alpha-3, alpha-4, and alpha-5, form a complex called a protein trimer. This trimer is a constituent factor of the basement membrane in the part of the kidney called the glomerulus. The glomerular basement membrane is a physical barrier against leakage of blood components during blood filtration, an important kidney function for producing urine. A mutation in any one of the three polypeptide chains can disrupt trimer formation and result in the development of AS due to a failure to properly form the basement membrane. Therefore, it is necessary to search for compounds that can correct or assist trimer formation by mutant collagen.
The researchers used split NanoLuciferase (split NanoLuc, NanoBiT) technology for protein-protein interaction analysis as a method of evaluating trimer formation. Fragments of two large and small luciferase molecules were fused with alpha-3 and alpha-5 chains and expressed in cells with alpha-4 chains. Measurable luminescence was detected only when alpha-3, alpha-4, and alpha-5 chains could form trimers.
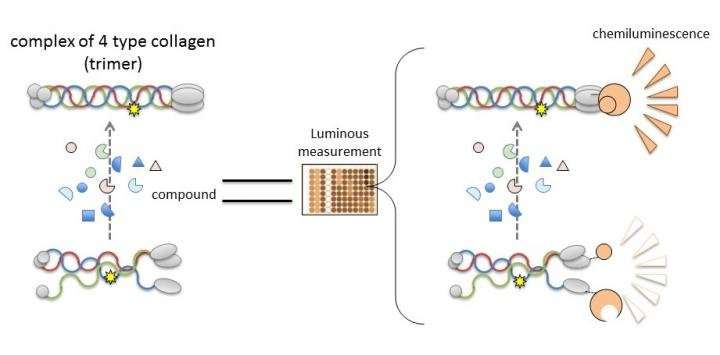
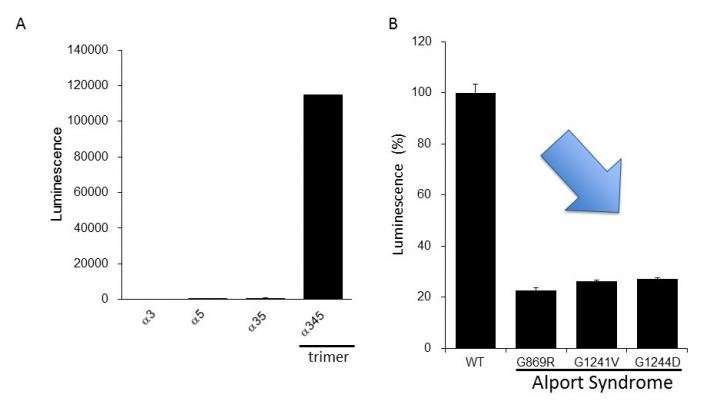
The researchers clarified that the NanoLuc-Col4 system reflects previously reported characteristics of Col4, namely that trimers are formed with a fixed combination of alpha-3, alpha-4, and alpha-5 chains, and that alpha-5 chains cannot form trimers if they are missing a functional part. The NanoLuc-Col4 system also revealed that alpha-5 chains with the gene mutations reported in AS failed to function. Importantly, as proof of principle that correcting trimer formation for mutant collagen is feasible, the authors were able to use the system to identify compounds that were able to induce trimer formation of alpha-3, alpha-4 and mutant alpha-5 collagen.
"This research should provide a pathway to develop drugs for hereditary diseases, such as Alport Syndrome, that are currently incurable," said Professor Hirofumi Kai of Kumamoto University's Department of Molecular Medicine.
More information: Kohei Omachi et al, A Split-Luciferase-Based Trimer Formation Assay as a High-throughput Screening Platform for Therapeutics in Alport Syndrome, Cell Chemical Biology (2018). DOI: 10.1016/j.chembiol.2018.02.003
Journal information: Cell Chemical Biology
Provided by Kumamoto University