A simple cell holds 42 million protein molecules, scientists reveal
It's official—there are some 42 million protein molecules in a simple cell, revealed a team of researchers led by Grant Brown, a biochemistry professor in the University of Toronto's Donnelly Centre for Cellular and Biomolecular Research. Analyzing data from almost two dozen large studies of protein abundance in yeast cells, the team was able to produce for the first time reliable estimates for the number of molecules for each protein, as revealed in a study published this week in the journal Cell Systems.
The work was done in collaboration with Anastasia Baryshnikova, a U of T alum and now Principal Investigator at Calico, a California biotechnology company that focuses on aging.
Proteins make up our cells and do most of the work in them. This way, they bring genetic code to life because the recipes for building proteins are stored within the genes' DNA code.
Explaining the work, Brown said that given that "the cell is the functional unit of biology, it's just a natural curiosity to want to know what's in there and how much of each kind."
Curiosity notwithstanding, there's another reason why scientists would want to tally up proteins. Many diseases are caused by either having too little or too much of a certain protein. The more scientists know about how protein abundance is controlled, the better they'll be able to fix it when it goes awry.
Although researchers have studied protein abundance for years, the findings were reported in arbitrary units, sowing confusion in the field and making it hard to compare data between different labs.
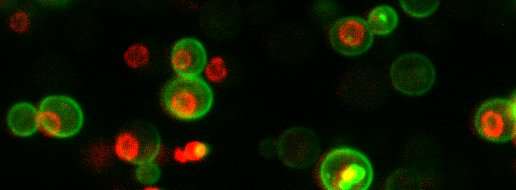
Many groups, for example, have estimated protein levels by sticking a fluorescent tag on protein molecules and inferring their abundance from how much the cells glow. But the inevitable differences in instrumentation meant that different labs recorded different levels of brightness emitted by the cells. Other labs measured proteins levels using completely different approaches.
"It was hard to conceptualize how many proteins there are in the cell because the data was reported on drastically different scales," said Brandon Ho, graduate student in the Brown lab who did most of the work on the project.
To convert arbitrary measures into the number of molecules per cell, Ho turned to baker's yeast, an easy to study single-cell microbe that offers a window into how a basic cell works. Yeasts are also the only organism for which there was enough data available to calculate molecule number for each of the 6,000 proteins encoded by the yeast genome thanks to 21 separate studies that measured abundance of all yeast proteins. No such datasets exist for human cells where each cell type contains only a subset of proteins encoded by the 20,000 human genes.
The wealth of existing yeast data meant that Ho could put it all together, benchmark it and convert the vague measures of protein abundance into "something that makes sense, in other words, molecules per cell," said Brown.
Ho's analysis reveals for the first time how many molecules of each protein there are in the cell, with a total number of molecules estimated to be around 42 million. The majority of proteins exist within a narrow range—between 1000 and 10,000 molecules. Some are outstandingly plentiful at more than half a million copies, while others exist in fewer than 10 molecules in a cell.
Analyzing the data, the researchers were able to glean insights into the mechanisms by which cells control abundance of distinct proteins, paving the way for similar studies in human cells that could help reveal molecular roots of disease. They also showed that a protein's supply correlates with its role in the cell, which means that it may be possible to use the abundance data to predict what proteins are doing.
Finally, in a finding that will rejoice cell biologists everywhere, Ho showed that the common practice of stitching glowing tags onto proteins has little effect on their abundance. While the approach has revolutionized the study of protein biology, netting its discoverers Osamu Shimomura, Martin Chalfie and Roger Tsien the Nobel prize in chemistry in 2008, it also stoked worries that tagging could affect protein durability, which would flaw the data.
"This study will be of great value to the entire yeast community and beyond," said Robert Nash, senior biocurator of the Saccharomyces Genome Database that will make the data available to researchers worldwide. He also added that by presenting protein abundance "in a common and intuitive format, the Brown lab has provided other researchers with the opportunity to reexamine this data and thereby facilitate study-to-study comparisons and hypothesis generation."
More information: Cell Systems (2018). www.cell.com/cell-systems/full … 2405-4712(17)30546-X
Journal information: Cell Systems
Provided by University of Toronto