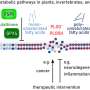
Study describes how the three-dimensional architecture of the genome changes during the cell cycle
Researchers at The Wistar Institute have uncovered new aspects of the three-dimensional organization of the genome, specifically how the genetic material is compacted and de-compacted in a timely fashion during the different phases of the cell cycle. This study was published in Nature Structural & Molecular Biology.
"We are just starting to appreciate that the way our genome is spatially organized in our cells has a profound impact on its function," said lead author of the study Ken-ichi Noma, Ph.D., associate professor in the Gene Expression and Regulation Program at Wistar. "Deciphering the three-dimensional structure of chromatin is essential to understanding key functions like transcription, DNA replication and repair."
The genetic information contained inside each of our cells is encoded by several feet worth of DNA molecules. Such an enormous amount of genetic material is packed into a microscopic space by folding it into a highly organized complex of DNA and proteins called chromatin. Although already significantly compacted, chromatin needs to be further condensed upon entry into mitosis, the process that divides each cell into two identical cells, in order to faithfully segregate the genetic material. Just as we pack up household belongings into smaller boxes when we relocate into a new house, it is easier to move and divide the DNA in the form of compacted chromosomes. This process has been known for several decades, yet the underlying molecular mechanisms that govern chromatin condensation and de-condensation are still poorly defined.
The Noma lab has extensively studied the mechanisms of genome organization using fission yeast as a model organism because it shares some important features with human cells while having a much smaller genome.
Noma and colleagues have previously described how two protein complexes called condensin and cohesin mediate the formation of functional genome-organizing structures called topological domains by establishing contacts that bring distantly located DNA regions closer together. Specifically, cohesin mediates local contacts, forming small topological chromatin domains, whereas condensin drives longer-range contacts, organizing larger domains.
In the new study, the lab applied similar genomic methodology to dissect the condensation and de-condensation of chromatin in topological domains over time, following the formation and decay of chromatin contacts throughout the different phases of the cell cycle. They discovered that the larger domains mediated by condensin are formed during mitosis, whereas the smaller, local domains mediated by cohesin remain stable throughout the whole cycle.
"Contrary to what was generally assumed in the field, we find that condensation and de-condensation of the chromatin domains happen very gradually and the cells oscillate smoothly between more and less condensed chromatin states," said first author of the study Hideki Tanizawa, Ph.D., an associate staff scientist in the Noma lab.
Alterations of the three-dimensional structures of the genome are linked to genetic diseases and cancer, presenting a powerful example of how basic cellular processes are relevant for disease. "The field is still in an early discovery phase but our study adds new insight into a fundamental biological process that may assist the development of novel therapeutic strategies in the future," added Noma.
More information: Architectural alterations of the fission yeast genome during the cell cycle, Nature Structural & Molecular Biology (2017). nature.com/articles/doi:10.1038/nsmb.3482
Journal information: Nature Structural & Molecular Biology
Provided by The Wistar Institute