Study suggests oysters offer hot spot for reducing nutrient pollution

When it comes to oysters and their role in reducing nutrient pollution, a new study by researchers at William & Mary's Virginia Institute of Marine Science gets right to the guts—and the shell—of the matter.
The study, in the September 29 issue of PLoS ONE, is the first to identify and quantify potentially denitrifying bacteria in the oyster gut and shell, and the first to do so using a new computer program that infers bacterial activities based on the sequences of ribosomal RNA genes.
Denitrification is the process by which nitrate and nitrite—compounds that fuel over-fertilization of coastal waters—are reduced to nitrogen gas, which is harmless to aquatic habitats. Excess nitrogen from wastewater treatment plants, farm fertilizers, and other human sources can lead to low-oxygen "dead zones," reduced fisheries harvests, and loss of sea-grass habitat. Chesapeake Bay is one of many ecosystems worldwide that have suffered these impacts.
Lead author on the study is Ann Arfken, a Ph.D. student in the lab of VIMS Associate Professor and co-author BK Song. Other co-authors are Drs. Jeff Bowman of the Scripps Institution of Oceanography and Michael Piehler of the University of North Carolina.
"Most studies addressing denitrification associated with oysters have focused on sediments in and around oyster reefs," says Arfken. "Ours is the first to explore the capability for denitrification by microbiomes living inside and on the oysters themselves." A "microbiome" is the community of microscopic organisms that inhabit each and every living thing, from the mites that dwell amid your eyelashes to the bacteria lodged among tomato roots.
The results of the study have important implications for efforts to reduce nutrient levels in coastal waters through oyster restoration. "We found that oyster shells contain unique microbial communities with higher denitrification activities than sediments," says Song. "It's thus possible to reduce nutrients at the beginning of an oyster restoration project as shell microbiomes are actively removing fixed nitrogen."
A financial and logistical challenge
Recent research shows that microbiomes play a key role in an organism's physiology and ecology, but, says Song, "Exploring the linkage between the genetic make-up and functions of a microbiome has long presented a challenge."
That challenge is both financial and logistical. "Studies of an organism's entire genetic make-up—its genome—are very expensive, and may not work for samples where the contribution from microbes is low," says Song. "As a result, many studies rely on amplicon sequencing of ribosomal RNA genes to identify microbial taxa." This approach is less expensive, but offers limited insight into the metabolic pathways associated with different microbiomes—in this case whether they possess the genes needed for denitrification.
To address these challenges, the team employed a new technique developed by Bowman. Called "PAPRICA"—for PAthway PRediction by phylogenetIC plAcement—it allows researchers to infer metabolic pathways from gene sequences associated with a small subunit of ribosome called16S rRNA.
"We combined a customized genomic database with the PAPRICA program to identify bacteria carrying denitrification genes among the microbiomes associated with oyster guts, shells, and reef sediments," says Song. He explains, "This would be somewhat like distinguishing among different human populations by comparing the number of stomach ailments with the gene needed to digest milk."
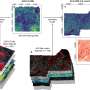
The research team then measured rates of denitrification in chambers containing live oysters, oyster shells, or sediments collected near oyster reefs, discovering much higher rates in the chambers holding living oysters and shells. When they compared these rates with the abundance of denitrification genes in the three microbiomes, they saw a strong correlation between high rates of denitrification and a gene sequence called nosZI.
"We found that bacteria carrying nosZI genes are important denitrifiers, and facilitate nitrogen removal in oyster reefs," says Arfken.
The researchers caution, however, that more research is needed. "We have to be careful when inferring denitrification and other metabolic processes from the presence of a gene in a bacterial genome," says Arfken. "These processes are often extremely complex, and require the coordinated expression of several different genes. Moreover, many organisms may carry a gene but not express it," A final challenge, she says, is that "lots of bacteria remain unclassified or have identified genomes that are either incomplete or of low quality."
But the researchers remain optimistic. "We're excited to pursue more studies to further validate the use of gene-based metabolic inferences as a reliable method for assessing the metabolic potential of microbiomes," says Song.
More information: Ann Arfken et al, Denitrification potential of the eastern oyster microbiome using a 16S rRNA gene based metabolic inference approach, PLOS ONE (2017). DOI: 10.1371/journal.pone.0185071
Journal information: PLoS ONE
Provided by Virginia Institute of Marine Science