The new method of analysis in record high speed DNA assay device

Molecular diagnostics plays an increasingly important role in medicine for the detection of genetic diseases, the monitoring of the effectiveness of anti-cancer therapy, and in the fight against aggressive bacterial infections. Curiosity Diagnostics (CD), a company belonging to the Scope Fluidics group, has developed a new technique for DNA analysis: synergistic PCR (sPCR). The method, described in detail in the joint publication of CD and IPC PAS researchers in Scientific Reports combines the advantages of two of today's most popular genetic code analysis techniques and can be carried out on widely available instruments.
"The DNA assay technique we propose was born during the development of the innovative PCR|ONE analytical instrument, which can be used to test the genetic code in only seven minutes. This is a more than ten-fold shorter time than is required in classic solutions," says Prof. Piotr Garstecki (IPC PAS, CD).
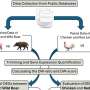
Samples forwarded for DNA assays usually contain so little genetic material that their analysis by conventional laboratory techniques would not be possible. After eliminating impurities from the sample, the first step is to increase the amount of genetic material, often by up to a billion times. polymerase chain reaction (PCR) is used for this purpose.
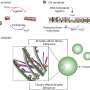
The PCR reaction involves the cyclical heating and cooling of a solution containing the genetic material being amplified and the appropriate reagents: a polymerase (i.e. the enzyme catalyzing the reaction of DNA synthesis), the nucleotides needed to build the DNA strand, and primers, i.e., short DNA fragments capable of attaching to the beginning and end of the propagated code fragment (e.g., a specific gene). Each PCR cycle consists of heating and cooling phases. In the first phase, at a temperature of about 95 degrees centigrade, the hydrogen bridges break, and the hitherto double-stranded DNA chain splits into two single strands. In the cool phase, at a temperature of about 50 degrees, the primers from the solution attach to the corresponding sites on the threads, after which the polymerase builds a complementary thread between them. At the end of each cycle, there are (in ideal conditions) twice as many double-stranded DNA fragments as at the beginning.
PCR is used to detect specific fragments of the genetic code and to estimate the original amount of genetic material. Quantitative measurements are usually carried out using an analogue technique known as real-time PCR. The sample is propagated, and in subsequent cycles, the amount of DNA is checked using fluorescent dyes. When the intensity of the changes exceeds a set threshold, the original amount of genetic material is estimated based on the number of cycles. The analogue PCR technique is relatively straightforward, but due to the sensitivity of PCR to even single particles of impurities, it requires careful, continuous calibration.
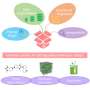
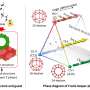
Another method is digital PCR. The sample is divided into tens or hundreds of thousands, and sometimes even millions of equal volume partitions. Then, in each partition, the procedure of duplication of genetic material is carried out and checks are made if the set change appears. During division of the sample DNA, molecules only reach some partitions, so the change does not occur everywhere. The original amount of DNA can therefore be estimated based on the number of recorded signals. The advantage of digital technology is that there is no need to calibrate the device. However, because of the need to conduct a large number of reactions in parallel, the testing equipment is expensive and is not as common in laboratories as analogue PCR apparatus.
Synergistic PCR, the technique proposed by CD and IPC PAS, combines the most important advantages of the analogue and digital methods. To obtain reliable measurements, it is enough to dilute a sample into only a dozen, or at most several dozen partitions. This method does not require calibration.
"A small number of partitions, characteristic of our technique, is of specific practical significance. It means that to perform the analysis, all that is needed is the standard well plate format used in popular analogue PCR devices," says Pawel Debski, an IPC PAS PhD student, who developed the sPCR method at Curiosity Diagnostics.
Due to a small number of sample partitions, the sPCR technique is easier to perform and slightly faster than digital variants. Compared to analogue techniques, however, more reagents are required. For this reason, it will not replace the analogue variant, according to the scientists. However, it may be a valuable addition because it needs no calibration, thus allowing laboratory staff to check the correctness of analogue measurements independently.
"Our DNA testing technique has been patented. However, we want to emphasize the freedom of using it for non-commercial purposes. And since it uses typical, popular genetic testing equipment, all you need do to get started is to reach for our article," highlights Prof. Garstecki.
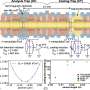
In standard PCR machines, relatively slow heat diffusion between the sample and an adjacent large block of alternately heated or cooled material is used to heat and cool the genetic material. In PCR|ONE, infrared radiation is used to heat the sample rapidly. The diffusion cooling mechanism has also been modified: the block used for this purpose is smaller than in conventional instruments and it is maintained at a constant, strictly controlled temperature. As a result of the technical and analytical improvements, the prototypes of PCR|ONE are able to complete DNA assays in less than a quarter of an hour, and the PCR itself takes only seven minutes. The first PCR|ONE devices will hit the market most likely in two to three years.
More information: Pawel R. Debski et al, Calibration-free assays on standard real-time PCR devices, Scientific Reports (2017). DOI: 10.1038/srep44854
Journal information: Scientific Reports
Provided by Polish Academy of Sciences