Rainbow dyes add greater precision to fight against 'superbugs'
A study reported Feb. 17 in the journal Science led by researchers at Indiana University and Harvard University is the first to reveal in extreme detail the operation of the biochemical clockwork that drives cellular division in bacteria.
The discovery, made possible through a revolutionary method used to color bacterial cell walls developed at IU, is an important step forward in research on bacterial growth and could inform efforts to develop drugs that combat antibiotic-resistant bacteria.
Globally, antibiotic-resistant bacteria, or "superbugs," pose a major risk to human health. The World Health Organization estimates about 480,000 people develop multi-drug resistant tuberculosis each year. In the U.S., the Centers for Disease Control estimates 1 in 4 hospital-acquired infections in long-term patients are caused by six major strains of the bugs.
"This is the first study to 'connect the dots' between each part of the cell involved in bacterial cellular division," said Yves Brun, the Clyde Culbertson Professor of Biology in the IU Bloomington College of Arts and Sciences' Department of Biology, who is an author on the study. "We've finally closed the circle on this mechanism and opened the door to more precise methods in the fight against antibiotic-resistant bacteria.
"If you understand how an engine works, you can shut it down by removing a single part," Brun said. "You no longer need to throw a hammer into the works to destroy it."
Early antibiotics like penicillin function like a hammer: a blunt instrument that destroys the bacterial cell in the midst of division by tricking cell wall-making enzymes called penicillin-binding proteins, or PBPs, into binding to the drug rather than the building blocks of the cell walls, causing the walls to breach and the cells to explode.
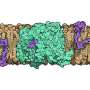
Other parts of the cell that drive bacterial division include cytoskeletal proteins, called FtsA and FtsZ, which form skeleton-like fibers inside cells to direct construction of the cell wall. All three elements must coordinate to build a cell wall in the middle of the cell to ensure the material inside doesn't escape after it splits in half.
The fact that these three parts of the cell play a role in cellular division is known, but the new study is the first to show exactly how they coordinate. Essentially, Brun said, FtsZ acts as a "foreman" that directs the movement of PBP "workers" as they construct a cell wall.
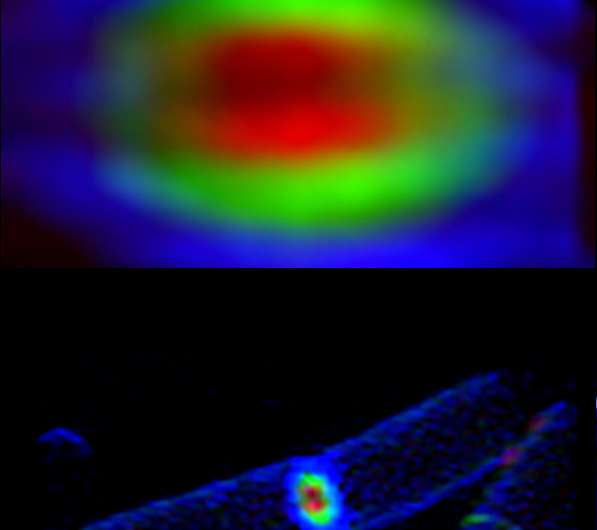
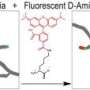
The researchers were able to detect the action with high-tech, multi-colored dyes called fluorescent D-amino acids, or FDAAs, discovered five years ago in the lab of Michael VanNieuwenhze, professor in the IU Bloomington College of Arts and Sciences' Department of Chemistry, who is a co-author on the study.
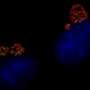
"The application of different colors of these dyes during the cell wall construction process revealed a 'bull's-eye pattern,' indicating the circular wall is built from the outer edge of the cell inward to the center," VanNieuwenhze said.
The study also solves another mystery: How do FtsZ molecules build the wall? The researchers found that FtsZ—which is arrayed in a biochemical chain called a filament—constantly loses a molecule at one end and gains a molecule at the other end, resulting in a circular motion around the cell's edge described as "treadmilling."
IU researchers chemically labeled the cells for analysis. Harvard scientists performed the experiments that showed the motion of the FtsZ and PBP proteins inside the cell.
The subject of a U.S. patent filed by the IU Research and Technology Corp., FDAA dyes have played an important part in dozens of other scientific papers on bacteria since 2012. VanNieuwenhze's lab also has about 50 material transfer agreements with researchers across the globe to provide access to the tool.
The creation of the dyes at IU was led by Erkin Kuru, a former Ph.D. student in the labs of VanNieuwenhze and Brun who is currently a research fellow at Harvard. Kuru and Yen-Pang Hsu, a IU Ph.D. student also in the labs of VanNieuwenhze and Brun, are co-authors on the study.
"This is the first time we've been able to observe cell division as a dynamic process—that is, a process occurring over time," Kuru said. "This wasn't possible before since we lacked the tools to see it."
Hsu added that "the visualization of these cell structures is no small task when you consider the organism that contains them is less than a micrometer—or one-thousand of a millimeter—wide. We wouldn't have been able to measure the fluorescent patterns in these cells without the technology at the IU Light Microscopy Imaging Center."
More information: "Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division" Science, science.sciencemag.org/cgi/doi … 1126/science.aak9973
Journal information: Science
Provided by Indiana University