Know thy enemy: Kill MRSA with tailored chemistry
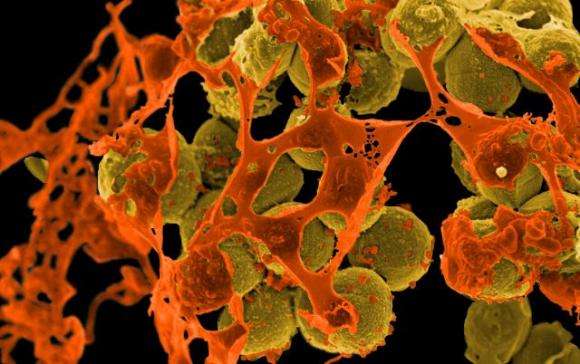
University of Connecticut medicinal chemists have developed experimental antibiotics that kill MRSA, a common and often deadly bacteria that causes skin, lung, and heart infections. The success is due to their strategy, which found a weakness and exploited it in a way the bacteria should have trouble countering, the researchers report in the Dec. 22 issue of Cell Chemical Biology.
Cases of MRSA (Methicillin-Resistant Staphylococcus aureus) are on the rise and increasingly resistant to common antibiotics. The first choice treatment for MRSA, trimethoprim-sulfamethoxazole, is relatively safe and inexpensive. But trimethoprim-resistant MRSA has begun to spread around the globe. Up to 30% of infections in sub-Saharan African no longer respond to it, and significant numbers in Europe and Asia as well. UConn medicinal chemists Amy Anderson, Dennis Wright and Ph.D. student Stephanie Reeve have been working to develop a drug that will be harder for MRSA to evolve resistance against. They had several candidates in the works when they asked colleagues at UConn Health and Hartford Hospital to start collecting trimethoprim-resistant strains of MRSA as test cases.
"Although resistance [to trimethoprim] in the community is generally less than 10% in our local area, resistance elsewhere is climbing. Additionally, many vulnerable patient populations cannot take trimethoprim-sulfamethoxazole or other generic drugs because of side effects they may cause, and new agents are needed," says Dr. Michael Nailor, a UConn pharmacologist co-funded with Hartford Hospital.
The local samples showed just how fast antibiotic resistance is spreading. Six of nine bacterial strains collected had genes for trimethoprim resistance that had never before been seen in the US. The strains were also variously resistant to other antibiotics such as erythromycin and tetracycline.
But they didn't stand a chance against the experimental antibiotics from Anderson, Wright and Reeve's lab.
"We've actually taken strains [of MRSA] from the clinic and shown our compounds work. We were really happy about these results," says Stephanie Reeve. Their strategy had worked.
"One of the most exciting aspect of this work was that we had worked hard to design broadly acting inhibitors against many different resistant forms of the enzymes and these designs proved very effective against two new enzymes we had never considered or previously studied," says Wright. The strategic approach the chemists had taken was to target the bacteria's use of Vitamin B9. Also known as folate, it's as critical to MRSA bacteria as it is to us. Block its action and a vital enzyme pathway is shut down. The bacteria die. Trimethoprim is currently the only antibacterial antifolate available, and bacteria have evolved different versions of the folate enzyme that aren't impaired by it. But Anderson, Wright and Reeve thought they should be able to make other, better antifolates. They painstakingly analyzed the molecular structure of the enzyme they were up against, and exactly how it needed to interact with other molecules to do its job. Only by understanding its form and function could they foil versions of the enzyme they'd never seen before, Wright says.
Armed with their knowledge, they designed new antifolates. These drugs are crafted to bind the enzyme in such a way that if the enzyme changes enough to evade them, it won't be able to do its job with vitamin B9, either. That will hopefully make it harder for bacteria to evolve resistance. The drugs' success against the trimethoprim-resistant strains of MRSA sampled so far bodes well.
The team is now gathering more MRSA from across the country. Dr. Jeffrey Aeschlimann, a UConn Health associate professor of pharmacy practice says, "We'd like to determine if the resistance mechanisms we discovered in our local MRSA strains are also found in other clinics throughout the United States. We also may find other novel resistance mechanisms. In both cases, we will be able to gain even more valuable information about our how our new antibiotics work against MRSA."
More information: Stephanie M. Reeve et al, MRSA Isolates from United States Hospitals Carry dfrG and dfrK Resistance Genes and Succumb to Propargyl-Linked Antifolates, Cell Chemical Biology (2016). DOI: 10.1016/j.chembiol.2016.11.007
Journal information: Cell Chemical Biology
Provided by University of Connecticut