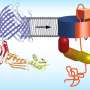
How drug-resistant bacteria build defences

Improved understanding of the way hundreds of different types of disease-causing bacteria operate could help pave the way to tackling their effects, according to leading scientists.
Researchers at the University of Leeds led by Professors Sheena Radford and Neil Ranson have uncovered new information about the operation of a protein complex called BAM – the beta-barrel assembly machinery – which helps insert other bacterial proteins into the bacteria's outermost protective layer.
In order for bacteria to spread diseases, they need to form an outer membrane, or they cannot survive in the harsh environments in which they live.
The BAM complex plays a crucial role in this process, facilitating the insertion of a myriad of proteins into this layer that are required for it to work properly as a barrier, protecting the bug from attack by antibiotics.
With their new results, researchers in the Astbury Centre for Structural Molecular Biology say it may be possible to design drugs which could target the BAM complex to stop it operating.
Professor Ranson said: "This is a complex molecular machine involved in keeping bacteria alive, and we know that mutations in it are lethal to the bacteria.
"Some beautiful work by other labs has shown that BAM exists in two different shapes – an open and closed form - but the open form seemed to require part of the complex to fall apart.
"Our work shows for the first time the intact BAM complex in this open state. Now we have been able to see the intact structure in this way, it gives us new clues about how BAM works and about how to develop ways to stop it functioning."
The research was carried out using electron microscopes based at the Diamond Light Source, the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
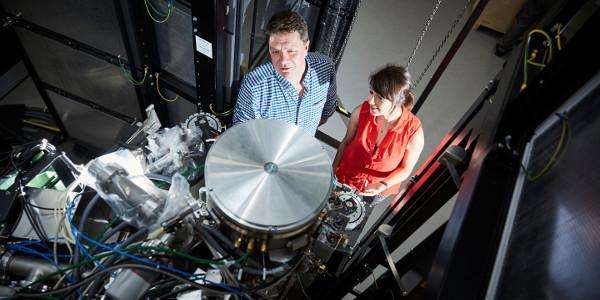

However, it is now possible to carry out the next steps in this research on campus in Leeds, following the University's investment in its own state of the art electron microscopes. The two microscopes form a key part of the leading edge facilities at the Astbury Centre for Structural Molecular Biology.
Professor Radford, Director of the University's Astbury Centre, said "If BAM cannot do its job, bacteria won't survive and they would be prevented from spreading diseases.
"The challenge, now we understand more about how BAM works, is for the scientific community to develop drugs which can target it and replenish our shrinking arsenal of antibiotics to treat bacterial infections."
The new understanding will contribute to ongoing work to find new ways to kill bacteria which have become increasingly resistant to drugs developed in 20th century.
Antibacterial resistance has become a major issue in recent years, with many drugs developed to combat infections from whooping cough to salmonella, and a wide range of hospital-acquired infections, losing their effectiveness.
Previously scientists have used X-ray crystallography, which has been the mainstay of structural discovery for several decades, to show that the barrel-shaped BAM complex could exist in both open and closed forms.
The BAM complex sits within the outer membrane of Gram-negative bacteria, and inserts other proteins into the membrane where they are needed to form a functional protective layer. How this happens is not properly understood, but the change in shape is thought to be essential to allow its client proteins to enter the membrane.
Using highly advanced electron microscopes, Professors Radford and Ranson now have seen the structure of BAM in an open state by rapidly freezing the complex from solution.
Previous structures of the open state all lacked one of the five proteins that make the full BAM complex, called BamB. The new results show that opening of the BAM barrel can happen with BamB there, and this changes the way scientists think about how BAM might work.
More information: Matthew G. Iadanza et al. Lateral opening in the intact β-barrel assembly machinery captured by cryo-EM, Nature Communications (2016). DOI: 10.1038/ncomms12865
Journal information: Nature Communications
Provided by University of Leeds