Scientists pave way for new generation of superbug drugs
Scientists at the University of East Anglia are getting closer to solving the problem of antibiotic resistance. New research published today in the journal Nature reveals the mechanism by which drug-resistant bacterial cells maintain a defensive barrier.
The findings pave the way for a new wave of drugs that kill superbugs by bringing down their defensive walls rather than attacking the bacteria itself. It means that in future, bacteria may not develop drug-resistance at all.
Unravelling this mechanism could also help scientists understand more about human cell dysfunctions linked to disorders such as diabetes, Parkinson's and other neurodegenerative diseases.
The team, supported by the Wellcome Trust, used Diamond Light Source, one of the world's most advanced scientific machines, to investigate a class of bacteria called 'Gram-negative bacteria'.
Diamond produces intense light 10 billion times brighter than the sun, allowing scientists to explore almost any material in atomic detail.
Gram-negative bacteria is particularly resistant to antibiotics because of its cells' impermeable lipid-based outer membrane.
This outer membrane acts as a defensive barrier against attacks from the human immune system and antibiotic drugs. It allows the pathogenic bacteria to survive, but removing this barrier causes the bacteria to become more vulnerable and die.
The research team previously found an 'Achilles heel' in this defensive barrier. But exactly how this defensive cell wall is built and maintained - the 'assembly machinery' - was unknown until now.
Lead researcher Prof Changjiang Dong, from UEA's Norwich Medical School, said: "Bacterial multi-drug resistance, also known as antibiotic resistance, is a global health challenge. Many current antibiotics are becoming useless, causing hundreds of thousands of deaths each year. The number of super-bugs is increasing at an unexpected rate.
"Gram-negative bacteria is one of the most difficult ones to control because it is so resistant to antibiotics.
"All Gram-negative bacteria have a defensive cell wall. Beta-barrel proteins form the gates of the cell wall for importing nutrition and secreting important biological molecules.
"The beta-barrel assembly machinery (BAM) is responsible for building the gates (beta-barrel proteins) in the cell wall.
"Stopping the beta-barrel assembly machine from building the gates in the cell wall cause the bacteria to die."
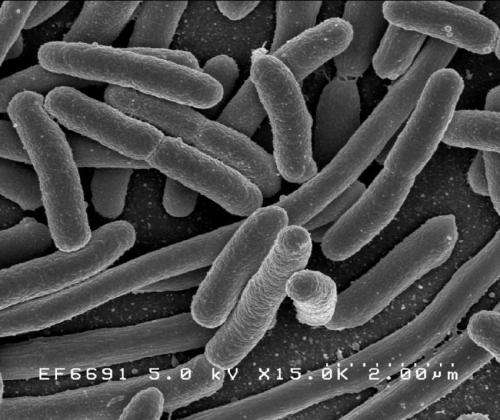
Scientists studied the gram-negative bacteria E.coli, in which the beta-barrel assembly machinery contains five subunits - known as BamA, BamB, BamC, BamD and BamE. They wanted to know exactly how these subunits work together to insert the outer membrane proteins into the outer membrane or cell wall.
Prof Dong said: "Our research shows the whole beta-barrel assembly machinery structures in two states - the starting and finishing states. We found that the five subunits form a ring structure and work together to perform outer membrane protein insertion using a novel rotation and insertion mechanism.
"Our work is the first to show the entire BAM complex. It paves the way for developing new-generation drugs.
"The beta-barrel assembly machinery is absolutely essential for Gram-negative bacteria to survive. The subunit BamA is located in the outer membrane and exposed to the outer side of the bacteria, which provides a great target for new drugs.
"In Human mitochondria, a similar complex called sorting and assembly machinery complex (SAM) is responsible for building the outer membrane proteins in the outer membrane of mitochondria.
"Dysfunction of mitochondria outer membrane proteins are linked to disorders such as diabetes, Parkinsons and other neurodegenerative diseases, so we hope that this work may also help us to better understand these human diseases too."
More information: 'Structural basis of outer membrane protein insertion by the BAM complex' is published in the journal Nature on February 22, 2016. DOI: 10.1038/nature17199
Journal information: Nature
Provided by University of East Anglia