Probing a mosquito protein for clues in the fight against Zika
As health departments around the U.S. boost efforts to combat Zika, scientists are working on new ways to kill the mosquitoes that carry the virus. One approach involves understanding the molecular mechanisms that keep the bugs alive so we can then undermine them. Scientists report in the ACS journal Biochemistry that they have revealed new structural insights on a key protein from Aedes aegypti, the mosquito species most often linked to the spread of Zika.
In February, the World Health Organization called for action against the disease after Brazil experienced a spike in the number of babies born with microcephaly, a condition characterized by an abnormally small head. Since then, the virus has been reported in more than 40 countries. Studies have shown that compounds that inhibit a protein called sterol carrier protein 2 (SCP2), which is involved in the transport of cholesterol and fats in insects, can kill Aedes aegypti larva. Kiran K. Singarapu and colleagues from CSIR - Indian Institute of Chemical Technology wanted to take a closer look at the structure of one of the protein's variants to help inform the development of future insecticides.
Using solution nuclear magnetic resonance, a technique that yields molecular-level information about proteins, the researchers were able to describe the 3-D structure and dynamics of a SCP2 variant. The new insights could help scientists screen small-molecule libraries for insecticide candidates. In addition to curbing Zika, any resulting compound that stamps out Aedes aegypti could reduce cases of other illnesses—dengue fever, yellow fever and chikungunya—that the mosquito also carries.
More information: Kiran Kumar Singarapu et al. Solution Nuclear Magnetic Resonance Studies of Sterol Carrier Protein 2 Like 2 (SCP2L2) Reveal the Insecticide Specific Structural Characteristics of SCP2 Proteins inMosquitoes, Biochemistry (2016). DOI: 10.1021/acs.biochem.6b00322
Abstract
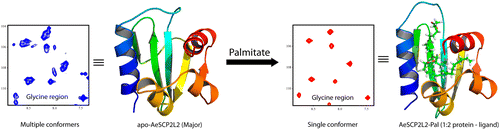
Sterol carrier protein 2 like 2 from Aedes aegypti (AeSCP2L2) plays an important role in lipid transport in mosquitoes for its routine metabolic processes. Repeated unsuccessful attempts to crystallize ligand free SCP2L2 prompted us to undertake nuclear magnetic resonance (NMR) spectroscopy to determine its three-dimensional structure. We report here the three-dimensional structures and dynamics of apo-AeSCP2L2 and its complex with palmitate. The 15N heteronuclear single-quantum coherence spectrum of apo-AeSCP2L2 displayed multiple peaks for some of the amide resonances, implying the presence of multiple conformations in solution, which are transformed to a single conformation upon formation of the complex with plamitate. The three-dimensional structures of apo-AeSCP2L2 and palmitated AeSCP2L2 reveal an α/β mixed fold, with five β-strands and four α-helices, very similar to the other SCP2 protein structures. Unlike the crystal structure of palmitated AeSCP2L2, both solution structures are monomeric. It is further confirmed by the rotational correlation times determined by NMR relaxation times (T1 and T2) of the amide protons. In addition, the palmitated AeSCP2L2 structure contains two palmitate ligands, bound in the binding pocket, unlike the three palmitates bound in the dimeric form of AeSCP2L2 in the crystals. The relaxation experiments revealed that complex formation significantly reduces the dynamics of the protein in solution.
Journal information: Biochemistry
Provided by American Chemical Society

















