Crowdsourcing the transformation of mass spectrometry big data into scientific living data
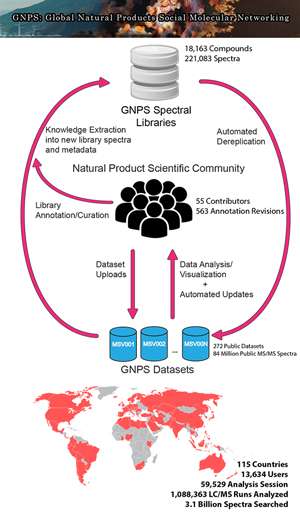
In a landmark paper published in the August issue of Nature Biotechnology, 127 scientists from a consortium of universities and research labs in the U.S. and worldwide report for the first time on the establishment of an online, crowdsourced knowledge base and workbench that could be a game-changer for the study of natural products that could potentially be useful in the development of the next antibiotic, better pesticides, or more effective cancer drugs.
"The potential of the diverse chemistries present in natural products remains untapped because natural product databases are not searchable with raw data and the research community has no way to share data other than through published papers," the paper noted. "Mass-spectrometry (MS) techniques are well-suited to high-throughput characterization of natural products, so there is a pressing need for an infrastructure to enable sharing and curation of the data."
Enter Global Natural Products Social Molecular Networking (GNPS), an open-access knowledge base for community-wide organization and sharing of raw, processed or identified tandem mass spectrometry data. The platform has operated in beta mode since 2014 under the leadership of the UC San Diego-based Center for Computational Mass Spectrometry (CCMS) and the Collaborative Mass Spectrometry Innovation Center in the Skaggs School of Pharmacy and Pharmaceutical Sciences, also at UC San Diego.
To date, more than 13,600 researchers from 115 countries have used the GNPS platform, utilizing various features and tools developed by CCMS to enhance or accelerate how researchers handle MS data (by allowing researchers to visualize/group the relationships of related molecules over petabyte-scale data spanning over one million samples). As the user community expands, so will its impact on science: so far, more than 100 different scientific papers have cited their use of the platform, with many different laboratories using it.
Since its launch, researchers have been developing the service to meet the needs of a broad population of scientists interested in natural products, and not just chemists. "We're facilitating making natural product chemical structures accessible to scientists who don't necessarily have training in chemistry," said Skaggs School of Pharmacy professor Pieter Dorrestein, one of the lead authors of the Nature Biotechnology paper. Dorrestein is also a key plater in the UC San Diego Center for Microbiome Innovation. "GNPS has created a community that is now willing to share knowledge, and we're enabling that community to do so, and this is unique in the natural products community."
To capture the knowledge of the community, the authors have created a crowdsourced, wiki-like knowledgebase that accepts submissions and revisions of annotated MS data from any scientist. Embracing a spirit of sharing and openness, these submissions are immediately and freely available to the entire community to be used to search MS data. "Just as BLAST finds regions of similarity between biological sequences in the form of nucleotide or protein sequences, the search function in GNPS serves a similar purpose," said CCMS Executive Director Nuno Bandeira, a UC San Diego professor with joint appointments in Computer Science and Engineering as well as the Skaggs School of Pharmacy, lead author of the Nature Biotechnology paper. "It allows researchers to find identical or similar molecular structures by comparing the chemistry based on spectral signatures captured using mass spectrometry."
Why the focus on natural products? Natural product scientists typically do not yet take full advantage of the modern capabilities of mass spectrometry.
"GNPS is the equivalent of BLAST, but for searching, analyzing and storing chemical signatures of molecules," said Dorrestein. "And like BLAST, GNPS is a tool that can be used by many different scientific communities."

Users at GNPS have analyzed more than one million LC/MS 'runs', with each instrument run typically processing approximately 3,000 MS/MS spectra. Correspondingly, the platform has processed more than 3.1 billion MS/MS spectra. An MS/MS search involves searching each MS spectrum (i.e., a molecular signature) for identical or similar MS spectra in the community database.
"At the beginning, people wondered if this would really be useful to them," said Mingxun Wang, a senior Ph.D. student in Computer Science and Engineering, who is the first author on the Nature Biotechnology paper. "Now, when we go to conferences, people come up and say, this is awesome! But they also make suggestions for new features, and we try to deliver them if we hear the same feedback from other users. One thing we did well is understand what is good for the community and a good user experience for our users, and that took on a life of its own."
One of the benefits offered by the platform is the ability to deliver frequent automated reanalysis of public data, taking into account the most recent user contributions to the MS knowledgebase. "We introduced the concept of 'living data' through continuous reanalysis of deposited data," said Bandeira. "We have this process of continuous identification, where users submit new annotations of molecules and all the data is re-searched against all the new annotations. We sometimes start with five or ten identifications, and after continuous searching we may typically now have hundreds of identifications. The knowledge keeps growing, and it keeps being part of the process."
Getting researchers to voluntarily contribute data to GNPS should benefit everyone because science depends on peer review to confirm results and develop new hypotheses. "The Big Data problem in biology cannot be solved in any single laboratory or one paper," said Bandeira. "The only way to extract value is through a community where everyone benefits from their contribution."
The scientists behind GNPS are among the first to recognize that GNPS is still in its infancy, but it also represents a huge investment in facilitating stepped-up research on natural products. "This resource is possible because CCMS was funded to build a platform to reach out to the community," said Bandeira. "Behind it are eight years of multiple people working out the system, which enabled us to implement all of these other features on top of it, including the ability to automatically schedule jobs to a cluster, a platform for web access, the forms and features; all of that had to be created from scratch before we could make these far-reaching contributions to the community as a whole."
In a 2014 paper, Dorrestein warned that fully integrated workflows to allow for rapid characterization involving millions of spectra "will only be possible if the community gets involved, shares and contributes data, and works to annotate the chemistry as a community." Now, says Dorrestein, that is happening. "GNPS is also changing the concept of how we interact with data," he says. "Usually scientists only deposit data if it's required, and after that, they don't care about it anymore. We want to change that paradigm. They share because this is the way to maximize their discovery potential. We make it easier for them to share their work with others. This should be seen across the board in all biology as a way to explore Big Data, and it's fantastic to do this in natural products, where this type of sharing could lead to the discovery of new antibiotics."
Building on a joint vision of the Bandeira and Dorrestein labs, Bandeira's CCMS was primarily responsible for developing the cyberinfrastructure and software tools to support GNPS, whereas Dorrestein's center contributed the data, reference collections, and many hours of personnel time on annotating data, and the researchers expect to continue the partnership between the data side and the software side. Added Bandeira: "The spirit of partnership and mutual benefit has been crucial to GNPS from the very beginning and continues to guide our vision for GNPS as a collaborative community resource aiming to help annotate all natural products MS data in the world."
That simple formula appears to be a hit with the user community. "We wanted to make it as simple as possible," said Dorrestein. "We created as few barriers as possible to make this happen, and that may be part of why people try it in the first place."
More information: Mingxun Wang et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking, Nature Biotechnology (2016). DOI: 10.1038/nbt.3597
Journal information: Nature Biotechnology
Provided by University of California - San Diego