Engineered gene drives and the future

Engineered gene drives, which have the potential to spread desirable genes throughout wild populations or to suppress harmful species, have received a lot of recent attention because of their potential to control organisms, such as mosquitoes that carry diseases such as Zika virus, malaria and dengue fever.
At the same time, the recently discovered CRISPR gene editing technology has the potential to create, streamline and improve the development of gene drives.
In a highly innovative and technical review, an entomologist at the University of California, Riverside has examined the different gene drives systems, analyzed the pros and cons of each and applications associated with them, and also surveyed the safety and regulatory issues associated with them.
"Despite all the potential benefits of gene drives, they remain understudied," said Omar Akbari, an assistant professor of entomology at UC Riverside. "With that in mind, and with advances occurring so quickly, we wanted to step back and take a broad look at what is happening."
Akbari, who is also a member of UC Riverside's Center for Disease Vector Research and Institute for Integrative Genome Biology, is the corresponding author of the piece, "Cheating evolution: engineering gene drives to manipulate the fate of wild populations," that was just published online in the journal Nature Review Genetics. The piece was co-authored by Jackson Champer and Anna Buchman, both post-doctoral students working with Akbari.
The idea of editing the genes of organisms to address biological problems related to public and environmental health has been around for decades. In fact, the authors cite a paper from 1940 and others from the late 1960s that discuss this.
Despite the wide-ranging applicability and importance of gene drives, there has been only modest development in their progress in past decades. Gene drives capable of functioning in wild populations have been created in only a few organisms, including yeast, the fruitfly and two species of mosquitoes.
This is, in part, due to the difficulty of engineering the genomes of organisms. However, recent advancements have provided tools capable of engineering the genomes of diverse species. The most promising of these tools is CRISPR.
Combining gene drives and a tool such as CRISPR may enable the development of novel strategies to reduce or eliminate insect-borne diseases, remove invasive foreign species, and even reverse the development of resistance to insecticides and herbicides, in an economically viable and environmentally friendly manner.
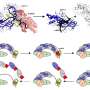
In the paper, the authors focus on several types of gene drives, including homing-based drives, sex-linked meiotic drives, medea and underdominance gene drives. They describe the gene drives based on different attributes, including rate of spread, species specificity, fitness cost, susceptibility to resistance, removability and reversibility.
They also discuss whether the gene drives are classified as 'modification drive' types, which are designed to spread through a population carrying desirable traits, or as 'suppression drive' types, which have the effect of reducing the population of a target species.
Finally, the authors address safety and regulation. They mention dangers associated with gene drives, including the potential to cause extinctions, to spread outside a geographical area or traverse into another species and the potential misuse to cause economic damage or even bioterrorism.
They write that the U.S. National Academy of Sciences has recently convened a panel to discuss the potential hazards and regulation of gene drives, and to make recommendations regarding their safe use. The authors say that there is no legislation specifically referring to gene drives and that their usage requires the need for local consent. They then add: "full transparency and early engagement with the public will be crucial for their approval."
More information: Jackson Champer et al. Cheating evolution: engineering gene drives to manipulate the fate of wild populations, Nature Reviews Genetics (2016). DOI: 10.1038/nrg.2015.34
Provided by University of California - Riverside