Researchers seek universally applicable technique for studying microbial metabolic activities

A team led by University of Vienna researchers has developed a way to identify and sort single microbial cells with the help of probes that use heavy water (laced with deuterium) and Raman microspectroscopy.
The approach is part of a universally applicable process being developed to characterize the activities of individual single cells. This allows researchers to better understand the roles microbes play within their native ecosystems.
As part of the 10-Year Strategic Vision of the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science User Facility at Lawrence Berkeley National Laboratory, the Emerging Technologies Opportunity Program (ETOP) was launched to support new technology development at other institutions that once perfected, can then be deployed by the DOE JGI and its users to tackle energy and environment applications.
One of these ETOP projects involves characterizing individual microbial cells in situ by combining labeling with heavy water, Raman microspectroscopy, microfluidics and flow cytometry. Proposed by researchers at MIT and the University of Vienna, Austria, the technology could accelerate the functional characterization of genes from metagenomic sequencing experiments, one of DOE JGI's highest priorities.
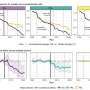
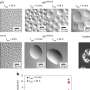
The early results of this ETOP project were described in the January 13, 2015 issue of the Proceedings of the National Academy of Sciences by a team led by University of Vienna researchers. Working toward a universally applicable technique that would allow researchers to evaluate microbial activities at the single cell level, the team conducted proof-of-principle experiments to study the efficacy of a process that involved incorporating heavy water (deuterium) isotopes into molecules, and then using Raman spectroscopy to chemically identify the microbes.
More information: "Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells." Proc Natl Acad Sci. 2015 Jan 13;112(2):E194-203. DOI: 10.1073/pnas.1420406112
Journal information: Proceedings of the National Academy of Sciences
Provided by DOE/Joint Genome Institute