Characterizing permafrost microbes in a changing climate

In the effort to curb climate change by reducing global greenhouse gas (GHG) emissions, thawing permafrost poses a critical challenge. These reservoirs of frozen organic matter embedded in Arctic soils are one of the major (~1.5 billion tons) stores of carbon on Earth. One of the abiding concerns regarding permafrost is that as global temperatures rise, as is projected over the coming centuries, soils may thaw completely. This event has the potential of causing the release of this carbon in the form of the potent greenhouse gases carbon dioxide (CO2) and methane, resulting in the largest contribution of carbon transferred to the atmosphere by a single terrestrial process.
To help understand the processes that control the conversion of organic matter to CO2 and methane, scientists from the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science user facility managed by the Lawrence Berkeley National Laboratory, reported on the application of multiple molecular technologies collectively referred to as "omics," to better characterize microbial activities in a paper published online March 4, 2015 in the journal Nature. The team also included members from Berkeley Lab's Earth Sciences Division, from Pacific Northwest National Laboratory (PNNL) and from the United States Geological Survey. The team sought to determine the composition of microbial communities and their role in degrading permafrost organic carbon and the subsequent production of CO2 and methane.
Microbial ecologist Janet Jansson from PNNL led the team that investigated three types of Alaskan soils, ranging from completely thawed to completely frozen. Metagenomics (MG), or environmental genomics, enabled the researchers to identify the phylogeny—i.e., history of organismal lineages—of the communities' microbial members, and the functional gene composition. Metatranscriptomics (MT) allowed the team to determine which genes were being expressed. Finally, metaproteomics (MP) provided insights on which proteins were actually produced.
Comparing microbial activities in various soils
The data collected for the permafrost study was impressive. "Together these analyses resulted in a large amount of data including 84.2 Gb [billion nucleotide bases] of MG sequence, 20.4 Gb of MT sequence, and approximately 7,000 proteins, which are among the highest yields obtained for any soil type to date," the team reported.
For the study, researchers relied on soil cores collected in Alaska, focusing on their bacteria and archaea. With the DOE JGI mobilizing its panoply of sequencing approaches and data analysis tools, the team studied these microbes in intact permafrost, the active layer above the frozen soil that seasonally freezes and thaws, and in other soils including the collapsed thermokarst bog that represents the terminal state of thaw.
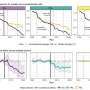
Comparison of the MG, MT and MP data from the three soils provided insight into the linkages between omics data and elemental cycling pathways. In the thermokarst bog they found the highest rates of methane production and identified several microbes involved in this pathway. Additionally, several genes involved in methanogenesis were detected in both the MG and MT data sets and corresponding proteins in the MP data sets. Three draft methanogen genomes were identified, and comparisons with sequenced methane producers suggest these are previously undescribed microbes.
The team found that the active layer had more diverse microbial species than the other soils, and the most active genes were involved in nitrogen, iron and methane cycles. In the permafrost samples, although fewer genes were expressed and fewer proteins were detected, they found that many of the proteins were those that allowed the microbes to tolerate the cold environment, more so than were found in the active layer and bog samples. They also found proteins in permafrost that suggest some of the microbes can move around, and proteins for pathways such as methane oxidation. The latter finding suggests that either the RNA and proteins have been preserved in the frozen environment, or the microbes with these functions are merely dormant in subzero conditions. Alternatively, they may represent physically active microbes in permafrost, thus providing a first insight into microbial survival strategies in permafrost.
Modeling microbial communities in Alaska
The work builds on findings from a previous collaboration between the DOE JGI and Jansson. In that study, which appeared in Nature, November 6, 2011, the draft genome of a novel methanogen was identified through environmental genomics studies that focused on Alaskan soil cores that contained both the active layer, and the permanently frozen soils beneath.
The work done by Jansson and her colleagues is just one of the ecosystem studies being conducted by the Department of Energy in Alaska. Through the Next-Generation Ecosystem Experiments (NGEE Arctic) project in Barrow, Alaska, a consortium of academic institutions and national laboratories is developing a process-driven ecosystem model that will allow researchers to better predict the evolution of Arctic ecosystems in a changing climate. Jansson is currently leading a Community Science Program project at the DOE JGI for sequencing of samples collected for the NGEE Arctic project.
The insights arising from this study are shedding light on microbial processes involved in GHG release, potentially as a consequence of temperature changes. A better understanding of these processes is necessary, the team maintained, for generating more accurate models and thus predictions of the environmental consequences.
More information: Multi-omics of permafrost, active layer and thermokarst bog soil microbiomes, Nature, DOI: 10.1038/nature14238
Journal information: Nature
Provided by DOE/Joint Genome Institute