New mechanism unlocked for evolution of green fluorescent protein

A primary challenge in the biosciences is to understand the way major evolutionary changes in nature are accomplished. Sometimes the route turns out to be very simple. An example of such simplicity is provided in a new publication by a group of ASU scientists.
They show, for the first time, that a hinge migration mechanism, driven solely by long-range dynamic motions, can be the key for evolution of a green-to-red photoconvertible phenotype in a green fluorescent protein (GFP).
Rebekka Wachter, a professor in ASU's Department of Chemistry and Biochemistry and the College of Liberal Arts and Sciences, is an expert in the field of structural characterization of GFP-like proteins. The present study is the culmination of eight years of intense effort in her laboratory.
The work, just published in the high impact journal Structure, involves collaborations with S. Banu Ozkan, from the Center for Biological Physics in the Department of Physics at ASU, and evolutionary biologist Mikhail Matz of the University of Texas.
Green fluorescent protein was first isolated from the jellyfish Aequorea victoria, which drifts with the currents off the west coast of North America. It was discovered that this protein glowed bright green under ultraviolet light. This phenomenon has found many creative applications in the biosciences.
The protein has been utilized as an extremely valuable luminous genetic tag for various biological phenomena. Using green fluorescent protein one can observe when proteins are made and where they go. This is done by joining the GFP gene to the gene of the protein of interest so that when the protein is made it will have GFP hanging off it. Since GFP fluoresces, one can shine light at the cell and wait for the distinctive green fluorescence associated with GFP to appear. The ability of some GFPs to turn red upon prolonged illumination makes them invaluable probes in super resolution fluorescence microscopy applications. This is where the current study is of most value.
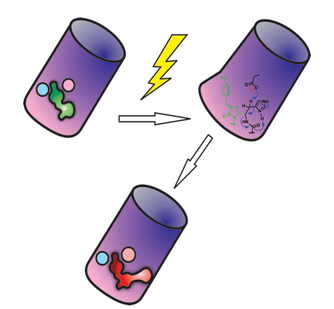
To fluoresce, GFP-like proteins must adopt a compact barrel-like shape. The light-triggered red phenotype may have arisen from a common green ancestor by a reversal of firm and soft regions located in opposite corners of the beta-barrel fold.
Although six crystal structures of reconstructed ancestral Kaede-type proteins indicate that the structure is highly conserved, analysis of chain flexibility by Molecular Dynamics and perturbation response scanning, performed in the group of S. Banu Ozkan has shown that the individual flexibility of the each position (i.e. structural dynamics) alters throughout the evolution of green-to-red photo conversion. Thus this study suggests that green-to-red photoconversion may have arisen from a common green ancestor by the shift of the rigid cornernear the chromophore to the opposite corner of beta-barrel.
"For the first time, this work establishes a direct experimental link between protein phenotypic change and collective dynamics without any external trigger, such as substrate, product or effector binding," explains Wachter. "Based on structural, computational and kinetic data, we propose a novel photoconversion mechanism that provides a plausible path for the irreversible acquisition of red fluorescence."
In spite of intense efforts in a number of laboratories worldwide, the mechanism of photoconversion of Kaede-type proteins has remained largely enigmatic. The present work sheds light on structural, dynamic and mechanistic features that must be considered when engineering improved fluorescent probes for super-resolution microscopy applications.
More information: Hanseong Kim, Taisong Zou, Chintan Modi, Katerina Dörner, Timothy J. Grunkemeyer, Liqing Chen, Raimund Fromme, Mikhail V. Matz, S. Banu Ozkan, Rebekka M. Wachter, "A Hinge Migration Mechanism Unlocks the Evolution of Green-to-Red Photoconversion in GFP-like Proteins," Structure, Volume 23, Issue 1, 6 January 2015, Pages 34-43, ISSN 0969-2126, dx.doi.org/10.1016/j.str.2014.11.011.
Journal information: Structure
Provided by Arizona State University