'Seeing' hydrogen atoms to unveil enzyme catalysis
(Phys.org) —Enzymes are catalysts that speed up chemical reactions in living organisms and control many cellular biological processes by converting a molecule, or substrate, into a product used by the cell. For scientists, understanding details of how enzymes work is essential to the discovery of drugs to cure diseases and treat disorders.
A multi-institutional research team led by Chris Dealwis from Case Western Reserve University has used the new IMAGINE instrument at Oak Ridge National Laboratory's (ORNL's) High Flux Isotope Reactor (HFIR) to map an enzyme that could play an important role in anti-cancer drug development.
Using neutron crystallography, the team unequivocally mapped the active site of the enzyme dihydrofolate reductase (DHFR), an enzyme necessary for nucleotide biosynthesis and a classical drug target. The team has been working toward unraveling the catalytic mechanism of DHFR for at least 15 years. The current research, published in the Proceedings of the National Academy of Sciences of the United States of America Early Edition in December, illustrates the pivotal role of neutron diffraction in unlocking the inner details of enzyme function by revealing protons in the enzyme-substrate, or Michaelis complex.
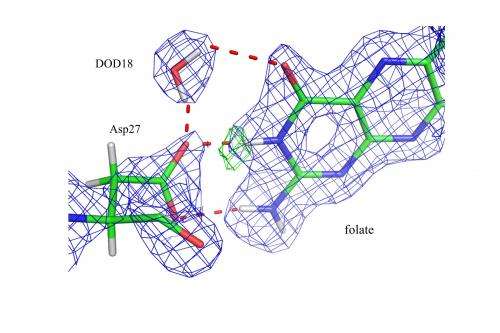
"We were able to detect protonation states at select sites in the enzyme to resolve mechanisms concerning acid-base catalysis," said Andrey Kovalevsky of ORNL, a co-author on the study and a scientist on the IMAGINE beam line at HFIR. The team essentially watched the enzyme work its magic, binding its substrate and preparing the active site for catalysis by protonating specific atoms and correctly orienting water molecules.
Using neutron diffraction, the team found that in a complex that closely resembles the reactive state, DHFR stabilizes a particular substrate's structure and likely elevates the basicity of the substrate atom that is to be protonated. This protonation is proposed to occur directly via water, with the proton's access to the substrate regulated by the structural fluctuation of the enzyme.
Because water, with its two hydrogen atoms, plays an important role in the chemical reactions catalyzed by enzymes, locating hydrogen atoms is necessary to map out the complete details of the enzymes' catalytic mechanisms. Hydrogen atom positions within an enzyme are difficult to pinpoint because they are hard to observe with traditional structural techniques; therefore, their positions are often inferred by those of nearby atoms.
"With neutrons, unlike other techniques like x-ray diffraction, we can truly see the hydrogen atoms and the orientation of the water molecules," said Kovalevsky. "Mapping enzymatic mechanisms is fundamental for our understanding of how nature works."
The enzyme DHFR is an anti-cancer target and is considered a model for understanding proton (H+) and hydride (H−) transfer reactions in biological systems. Given its role in biosynthesis, DHFR is a target for anticancer, antimicrobial, and rheumatoid arthritis drugs, such as methotrexate and trimethoprim.
This is the first user publication from the quasi-Laue diffractometer instrument IMAGINE at HFIR, beam line CG-4D. IMAGINE is a state-of-the-art neutron single crystal diffractometer using image-plate technology. It provides atomic resolution information on chemical, organic, metallo-organic and protein single crystals that will enable their chemical, physical and biological structure and function to be understood.
"We need more experiments and different samples to figure out how powerful the instrument is—how small its samples can be, how large the crystal unit cells we can resolve," said Kovalevsky. "But this result shows the instrument is performing to its specifications, which is exciting."
For future experiments, the IMAGINE team plans to introduce a low-temperature sample environment to collect data at cryogenic temperatures. Kovalevsky explains that they sometimes need crystals to be very cold to trap short-lived intermediates of enzymatic reactions, and to improve diffraction quality so that smaller crystals can be used for data collection.
More information: Q. Wan, B. C. Bennett, M. A. Wilson, A. Kovalevsky, P. Langan, E. E. Howell, and C. Dealwis. "Towards resolving the catalytic mechanism of dihydrofolate reductase using neutron and ultrahigh resolution X-ray crystallography." Proceedings of the National Academy of Sciences Early Edition. (2014). www.pnas.org/cgi/doi/10.1073/pnas.1415856111
Journal information: Proceedings of the National Academy of Sciences
Provided by Oak Ridge National Laboratory