Scientists manipulate split proteins to detect interactions in living cells

(Phys.org) —Rice University scientists have developed a plug-and-play approach to detect interactions between proteins they say could greatly improve understanding of basic biological functions.
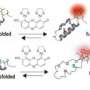
The Rice group of synthetic biologist Jonathan (Joff) Silberg, in collaboration with Baylor College of Medicine, split and added sticky tags to fluorescent proteins that become biosensors when inserted into living cells. When the sensors detect their target interactions, they reconnect and glow under near-infrared light.
The customizable technique could become a valuable toolkit for those who study protein interactions responsible for diseases, including cancer, and other biological functions that are difficult to analyze, Silberg said.
The research is the subject of a new paper in the American Chemical Society journal ACS Synthetic Biology.
Proteins are the molecules that carry out tasks in biological systems, either on their own or as part of signaling pathways that could involve many steps. Understanding how they work and why they sometimes break down is key to treating disease, Silberg said.
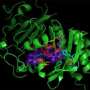
The Rice lab embarked on the project based on work by Baylor microbiologist Anthony Maresso, a co-author of the new paper who had been using a near-infrared fluorescent protein (IFP) to see deep into the tissues of living mice. IFPs are biologically inert proteins whose sole function is to fluoresce under near-infrared light.
"We got interested in this because we want to know what promoters, or switches, are turned on in a microbe's genes," Silberg said. "I want to know not just if one protein is being made but if two proteins are being made, and until now there's been no way to do that.
"So when Anthony told me his infrared fluorescent protein works in bacteria, I thought, 'If we break it into two pieces, then we could use two inputs to switch the light 'on' in this protein and make it glow."
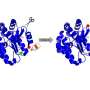
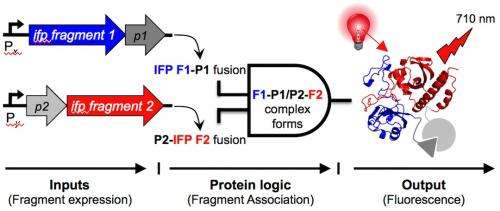
Silberg and lead author Naresh Pandey, a graduate student in his lab, discovered that even if they split the protein at random places, when they added peptide tags – fragments of proteins – to the newly created ends, the protein would reassemble and light up when it encountered its target. The tags determine the IFP's target and allow it to be customized for specific interactions.
"We had made every possible two-piece protein by splitting the IFP in different places, and we found that nothing lit up," Silberg said. "When we decided to give it a little help and added Velcro-like peptides to the ends, we found many two-piece proteins that glowed.
"That's what's exciting for us: Everything we found, all of these different split variants, were plug and play. We could plug in different protein-protein interactions and they still gave us a signal," he said.
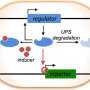
Through directed evolution, a way to create new proteins by mimicking natural selection, the Rice team made entire libraries of mutant IFPs that function as "AND" logic gates. These gates, like their counterparts in computer circuits, require the presence of one protein trigger and another protein trigger to come together and glow when hit with near-infrared light.
The IFPs are far better for deep biological studies than standard green fluorescent proteins (GFPs), Silberg said. "The light needed to turn on a GFP and make it glow doesn't get very far," he said. "It doesn't penetrate very well into deep tissue, because there are a lot of things that absorb light in that region of the spectrum. But near-IR offers us the best signal we can get with any protein."
Silberg said the system will be useful not only for medical researchers like Maresso who are interested in studying microbial pathogens, but also for those who study gene therapy technologies in animals, including co-author Lynn Zechiedrich, a Baylor College of Medicine professor who has pioneered the development of DNA MiniVectors.
"MiniVectors are small circular pieces of DNA that can get into many cells. However, they display the best DNA delivery properties at small sizes that cannot encode a full protein like IFP," Silberg said.
"What Lynn really needs are proteins that can be delivered to cells in pieces. Our two-piece IFP will enable her to study MiniVectors delivery of split-protein cargos in animals, and our approach for breaking proteins into pieces will allow us to build new therapeutic cargos that can be delivered to diseased cells using MiniVectors," he said.
Silberg suggested split IFPs could have many other uses. "What if you can fuse this protein to another that senses a chemical? Maybe a toxin or a metabolite? This opens up the possibility of engineering new complex biosensors for use in animals."
More information: "Combining random gene fission and rational gene fusion to discover near-infrared fluorescent protein fragments that report on protein-protein interactions." Naresh Pandey , Christopher L Nobles , Lynn Zechiedrich , Anthony W. Maresso , and Jonathan J. Silberg. ACS Synth. Biol., Just Accepted Manuscript, September 29, 2014. DOI: 10.1021/sb5002938
Journal information: ACS Synthetic Biology
Provided by Rice University