Attacking persister cells that are responsible for making bacteria resistant to new drugs
(Phys.org) —We face an urgent global health problem because scientists are not developing new antibiotics as fast as bacteria are developing antibiotic resistance.
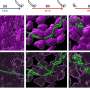
But new research from UCLA has made important progress toward solving this problem. An interdisciplinary team of scientists from UCLA's California NanoSystems Institute has developed a method to re-engineer antibiotics that sharply enhances their activity against certain key bacterial cells, called persisters, that are responsible for making bacteria resistant to new drugs.
Persister cells slow down their metabolism and shut down their mechanisms for taking in molecules, preventing normal antibiotics from getting into them, which is necessary for the drug to kill the bug. After the persister cells survive the initial antibiotic treatment, they pass on their genes as the bacteria reproduce.
Led by Gerard Wong, professor in the UCLA Department of Chemistry and Biochemistry and the Department of Bioengineering, and Andrea Kasko, associate professor of bioengineering, the team has developed a method analogous to taking an ordinary car and adding high-performance parts to make a fast and furious street racer.
"We're in an unsustainable race with bacteria. They become resistant to our antimicrobials too fast," Wong said. "It takes upwards of $100 million to develop one antibiotic drug, and bacteria develop resistance to it within two years. It's a race that we can't win. This reality brought us to the idea of taking an existing antibiotic and renovating it, giving it a new, complementary antimicrobial ability while preserving its original ability to make a better drug overall."
The study was published Aug. 18 online in the journal ACS Nano.
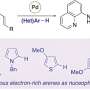
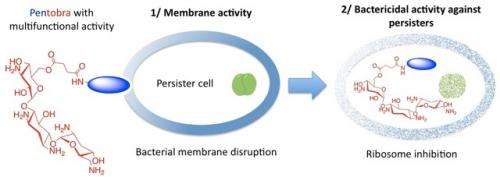
The team began with an antibiotic that many persisters are resistant to called tobramycin. By adding amino acids to the molecule of tobramycin, they created a new antibiotic drug molecule. The new compound, Pentobra, is able to easily punch holes in persister cell membranes and get inside the cells to kill the bacteria.
"What we've done is make a molecule that kills with more than one mechanism," Wong said. "Pentobra can punch enough holes in the cell membrane to kill the cell, but that may not be the most efficient way to kill a bug. This antibiotic also messes up their ability to grow by preventing them from making more bacterial proteins."
Wong said that the combination of sabotaging protein synthesis and punching holes in cell membranes is synergistic.
"The two mechanisms work with one another quite nicely, which might be what makes this antibiotic so powerful and why Pentobra can kill persister cell strains 10,000 to 1 million times better than tobramycin," Wong said. "Andrea and I thought it was a good idea to combine these two properties in engineering a drug molecule for persisters, but we were not expecting enhancement to that degree."
Kasko pointed out that the challenge of their design was that they had to make sure the combined elements of Pentobra would work well together and not deactivate each other, so that they could work synergistically against the target persisters.
"We had to make sure that the new drug molecule would still permeate the cell membrane while maintaining its ability to kill the cell once it was inside. Without properly designing the new molecule from the parent drug before we tried combining the components, we could have ended up with a completely inactive molecule," Kasko said.
This revolutionary process can be used to renovate other existing antibiotics to resurrect their activity against resistant bacteria and to enhance their potency. The new approach can also potentially be used to improve many new antibiotics being developed. According to Kasko, "If someone has a drug they think has potential, with our method they may be able to make it even more potent."
More information: "Engineering Persister-Specific Antibiotics with Synergistic Antimicrobial Functions" Nathan W. Schmidt , Stephanie Deshayes , Sinead Hawker , Alyssa Blacker , Andrea M. Kasko, and Gerard C. L. Wong. ACS Nano, 2014, 8 (9), pp 8786–8793 DOI: 10.1021/nn502201a
Journal information: ACS Nano
Provided by University of California, Los Angeles