Detecting tumor markers easily

Blood is just teeming with proteins. It's not easy there to identify specialized tumor markers indicating the presence of cancer. A new method now enables diagnostics to be carried out in a single step. Scientists will present the analysis equipment at analytica, the international trade fair in Munich April 1-4.
Benign growth, or cancer? Tumor markers in the blood help determine whether the patient is afflicted with a malign tumor and whether it is excreting markers more vigorously – involving highly specific proteins. An increased concentration in the blood provides one indication of the disease for physicians. However, it has been quite expensive in time and effort to detect the markers thus far. This is because all kinds of molecules and proteins are teeming in the blood. To be able to detect a single specific one, doctors must first separate and purify the blood in several steps, and then isolate the marker they are searching for from the rest of the molecules.
This will be faster in the future. Researchers in the Project Group for Automation in Medicine and Biotechnology PAMB of the Fraunhofer Institute for Manufacturing Engineering and Automation IPA in Mannheim, Germany, have developed a one-step analysis. "Our goal is to detect biological molecules in blood, or in other kinds of samples from the patient such as urine, that indicate diseases," explains Caroline Siegert, a scientist at IPA, "and do so without having to laboriously process the blood, but in one single step instead."
Lower noise, higher signal
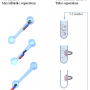
The difficulty in detecting specific molecules in the blood or urine lies in the enormous number of substances that are mixed in the liquid. They cause a high level of background noise that masks the desired signal. The signal from the protein being searched for can no longer be distinguished – similar to how a single voice in a room full of chattering people can be difficult to make out. "We are able to improve the signal-to-noise ratio using our technique, so we can recognize individual molecules even in blood samples," says Siegert. The researchers use magnetic beads for this – particles only a few micrometers in size that have a magnetic core. If you position a magnet externally on the test tube, you can arrest or steer the beads with it. This technique is already in use. To isolate molecules from a solution like blood, researchers coat the surface of the beads with specialized antibodies. If the proteins that are being searched for meander past the beads, the antibodies grab hold of them. If you now hold a magnet to the outside of the test tube, the beads together with the desired proteins stick to the interior surface of the tube, while the rest of the solution can be easily removed.
Synchronous glow
To determine immediately whether the hunt for the proteins has been successful the Fraunhofer researchers developed an additional means. The samples and investigation are not just exposed to the coated beads, but also to additional antibodies that have fluorescent markers bound to them. These antibodies attach themselves to the proteins being searched for and cause them to glow. The optical signal is very weak though, and would normally disappear in the background noise. However, a trick can be used to amplify it. If you apply an alternating magnetic field, the magnetic beads flock together in rhythm, and the fluorescent markers bound to the surfaces emit their light in sync – and radiate considerably more brightly than any given bead by itself. The advantage of this visualization technique is that the optical signal provides immediate insight about whether a tumor marker protein is present in the blood, for example. Long waits for expensive laboratory investigations are no longer necessary.
There are many applications for the new procedure. Not only can numerous biological molecules such as antibodies be detected, but also nucleic acids that indicate an infection. "Since we use conventional beads, it means the molecules can be quickly and simply isolated after the analysis as well," Siegart says.
Provided by Fraunhofer-Gesellschaft