Hugging hemes help electrons hop
(Phys.org) —Researchers simulating how certain bacteria run electrical current through tiny molecular wires have discovered a secret Nature uses for electron travel. The results are key to understanding how the bacteria do chemistry in the ground, and will help researchers use them in microbial fuel cells, batteries, or for turning waste into electricity.
Within the bacteria's protein-based wire, molecular groups called hemes communicate with each other to allow electrons to hop along the chain like stepping stones. The researchers found that evolution has set the protein up so that, generally, when the electron's drive to hop is high, the heme stepping stones are less tightly connected, like being farther apart; when the drive to hop is low, the hemes are more closely connected, like being closer together. The outcome is an even electron flow along the wire.
This is the first time scientists have seen this evolutionary design principle for electron transport, the researchers reported Jan. 2 in Proceedings of the National Academy of Sciences Early Edition Online.
"We were perplexed at how weak the thermodynamic driving force was between some of these hemes," said geochemist Kevin Rosso of the Department of Energy's Pacific Northwest National Laboratory. "But it turns out those pairs of hemes are essentially hugging each other. When the driving force is strong between hemes, they are only shaking hands. We've never seen this compensation scheme before, but it seems that the purpose is to allow the protein to transfer electrons with a steady flow along heme wires."
Living Wires
Certain bacteria breathe using metal like people use oxygen. In the process, these bacteria steal electrons from carbon and ultimately transfer the electrons to metals or minerals in the ground. They do this by conducting electricity along molecular wires built into proteins, moving internal electrons to the outside of their cells. Researchers hope to use these bacteria in little biologic batteries or fuel cells.
But a living wire is not the same as those that make up our powerlines. Electrons in powerlines hurtle down the wire, moving smoothly from metal atom to metal atom. Electrons traveling in a living wire must get from one complex heme group to the next. The hemes are situated within a protein, and not all hemes are made the same.
Some hemes hold onto electrons tightly and others let electrons slip away easily. Depending on how the hemes are lined up, this can create energetic hills that electrons have a hard time climbing over, or energetic valleys that electrons easily march across.
Some hemes, such as those that carry oxygen in people's red blood cells, are well-studied. The hemes and proteins creating a current in bacteria, though, have only been coming to light within the last few years. Recently, researchers figured out what a particular protein—MtrF—that makes up a molecular wire looks like, but that information alone is not enough to determine how the electrons traverse the chain of internal heme groups.
So, armed with the structure of the protein, Rosso and colleagues Jochen Blumberger and Marian Breuer from the University College London used high-powered computers to simulate the positions and movement of the hemes in MtrF and how they transfer electrons between themselves.
Electron Crossroads
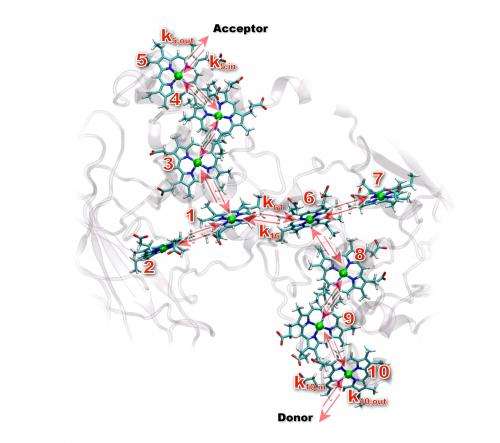
Using resources at both the UK's High Performance Computing Facility and EMSL, the Environmental Molecular Sciences Laboratory at PNNL, the team first modeled the average position of the 10 hemes within MtrF. Eight of the hemes run down the center of the protein. The remaining two hemes branch off the main eight, creating a four-heme road that crosses the middle of the protein.
Because hemes have to pass electrons to each other, the team examined them in pairs. The team found that MtrF arranges its heme pairs in one of three ways: perpendicular to each other, side-by-side, or stacked on top of each other. Each arrangement positions the hemes at different distances from and orientations to each other.
Then the team gauged how urgently an electron wants to get from one heme to the next by determining the theoretical "Gibbs free energy" between the pairs. This value is an indicator of the driving force of the electrons.
The team found that instead of a smooth ride through the protein, electrons lurch through hemes: Sometimes the driving force makes the electrons march across a valley and the electrons move quickly. In other pairs the electrons face a hill, and electron travel gets delayed.
Mapping how tightly hemes couple to each other along with the driving force values, the team found that hemes were less tightly coupled when electrons enjoyed traipsing across a valley and more tightly coupled when electrons had to slog uphill.
"The computer simulations allowed us to break the wire down into how each step is possible and how fast each step is. Then we saw that the protein arranges its hemes in weak and strong couplings to compensate for the energetic hills and valleys," said Rosso. "This is one way to make the electron hops consistent to efficiently get them where they need to go."
This compensation scheme led the team to wonder why the hills and valleys are there in the first place.
"We think the variation in driving force between the hills and the valleys helps the protein interact with other components in the environment," said Rosso. The tops of the hills could be exit points to higher energy electron acceptors in the environment, such as molecules that shuttle electrons elsewhere.
Scientists don't yet know how multiple heme proteins—including others beyond MtrF – work in concert to make these molecular wires connect end-to-end, but the results give hints as to which hemes are possible entry and exit points in MtrF. So the results also give clues to how multiple proteins might be connected.
More information: Marian Breuer, Kevin M. Rosso, and Jochen Blumberger. Electron flow in multiheme bacterial cytochromes is a balancing act between heme electronic interaction and redox potentials, Proc Natl Acad Sci, Early Edition online January 2, 2014. DOI: 10.1073/pnas.1316156111.
Journal information: Proceedings of the National Academy of Sciences
Provided by Pacific Northwest National Laboratory