Mother nature as a wire manufacturer: Scientists see how microbe directs electrons
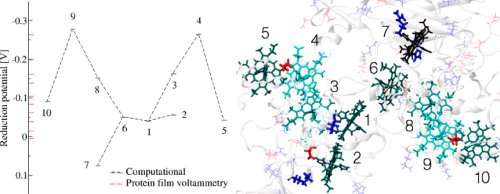
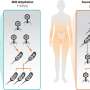
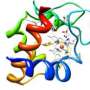
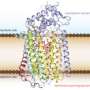
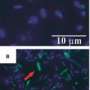
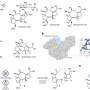
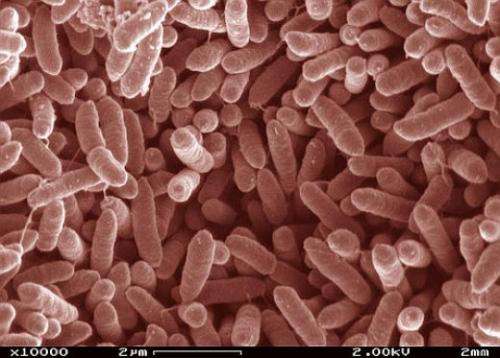
(Phys.org) -- For the first time, each step an electron takes as it moves along a "wire" from a microbe's interior to the outside world is known, thanks to modeling by University College London and Pacific Northwest National Laboratory. The wire is a twisted molecule, known as a cytochrome. The cytochrome is made of 10 iron-based clusters, called hemes, held in a particular arrangement by a protein that serves as a backbone. The hemes are positioned end-to-end in a staggered cross, to comprise the electron transfer interface between the interior of Shewanella oneidensis, a bacteria found in the soil, and metals and minerals beyond the cell's wall.
"This is the first look at how nature would design a molecular wire for transport on nanometer length scales," said Dr. Kevin Rosso, a geochemist at PNNL and a corresponding author on the study.
To understand how these microbes influence contaminated water and soil, as well as to answer the clamor for smaller and smaller computer circuits, scientists are looking to bacteria. Optimized over eons, bacteria are very proficient at moving electrons using molecular complexes. This study provides details about the molecules that move the electrons and how they work. It is opening doors to designing molecule-sized circuits and to research related to how these microbes affect contaminated water and soil.
Last year, the structure and other characteristics of the 10 hemes in the bacterial protein were established from experiment. The results appeared in Proceedings of the National Academy of Sciences. This year, Rosso and team wanted to determine exactly what was happening with the hemes as the electrons jump from one to the next, so they turned to computational modeling and simulation.
"Lab experiments can't go after that type of information," said Rosso. "From an experimental standpoint, the hemes all look alike; they can't isolate and identify one heme at a time. We can."
The team built large-scale computational simulations of the cytochrome that involved tens of thousands of atoms, allowing them to determine how easily the electrons were transferred heme to heme. Specifically, they wanted to know the redox potential of each iron-based complex, with the surrounding molecular environment and dynamics explicitly described. The redox potential is a measure of a molecule's tendency to acquire electrons. It turns out that the cytochrome's hemes are relatively balanced in their ability to pull electrons from each other, meaning that electrons do not strongly favor one direction over another. Further, under the right conditions, the protein looks able to turn the flow of electrons on and off, and might even be able to reverse the flow of electrons, based on subtle distortions in the molecular structure.
To verify the results, the simulations were run on two different supercomputers and then compared. The results agreed well and were recently published as a rapid communication in the Journal of the American Chemical Society.
Having determined the structure and how the electrons hopscotch from heme to heme, the researchers are now studying electron flow over longer time scales. Further, they are studying at how changes in the cytochrome might alter the electron's passage.
More information: M Breuer, et al. 2012. "Thermodynamics of Electron Flow in the Bacterial Deca-heme Cytochrome MtrF." Journal of the American Chemical Society 134(24):9868-9871. DOI: 10.1021/ja3027696
Journal information: Proceedings of the National Academy of Sciences , Journal of the American Chemical Society
Provided by Pacific Northwest National Laboratory