Molecular motors involved in chromosome transport observed
Researchers at Waseda University in Japan have for the first time directly observed the "molecular motor", called Xkid, that plays a critical role in facilitating the proper alignment of chromosomes during cell division. Their findings are expected to contribute greatly to elucidating the molecular mechanisms of chromosome segregation, a key aspect of the development of certain medical disorders including cancer and birth defects.
A human being begins as a single fertilized egg cell, which keeps dividing to form muscles, internal organs, the brain and other structures. At every step of cell division, chromosomes – the source of all genetic information – must be precisely segregated without any misplacement between two daughter cells. Otherwise, their incorrect segregation can cause a variety of medical disorders.
Within each cell, Xkid molecules are located inside the spindle apparatus, a structure required for cell division that's composed of a bundle of microtubules. Determining the movements of Xkid in natural spindles is considered the key to understanding the mechanisms of chromosomal segregation during cell division.
In a recent study published in the online journal Scientific Reports, Jun Takagi and colleagues at Waseda University reported the first molecular-level demonstration of how Xkid directs chromosomes toward the centre of the spindle apparatus. The study showed that Xkid molecules travelled along spindle microtubules, while the distribution of spindle microtubule polarity was optimal for Xkid-facilitated chromosome transport. This study provides invaluable knowledge in exploring the mechanisms of materials transport in biological systems.
While a human body is composed of many different parts such as muscles, internal organs, and a brain, its origin is only a single cell, a fertilized egg, which keeps dividing to form the human being. At every step of the cell division, chromosomes must be precisely segregated without any kind of misplacement between two daughter cells. Chromosomes are the source of all genetic information, and incorrect chromosome segregation can cause various forms of medical disorders including severe illnesses and malignant transformation of tumors.
The Xkid molecular motor is known to play a critical role in facilitating the proper alignment of chromosomes during cell division. Previously, its motor functions have been investigated in vitro using purified Xkid molecules obtained from cell extracts, showing their plus end-directed movement as ensembles of molecules along the microtubules. How such molecular motors behave within intact spindles, however, remained to be characterized. In a cell, Xkid molecules are located within the spindle apparatus, a structure required for cell division. The spindle apparatus is an orderly structure composed of a bundle of numerous microtubules. Elucidating Xkid movements in natural spindles is the key to understanding the mechanisms of chromosome segregation during cell division.
Accordingly, the present study set out to examine the movements of Xkid within an intact spindle.
Techniques developed during the study
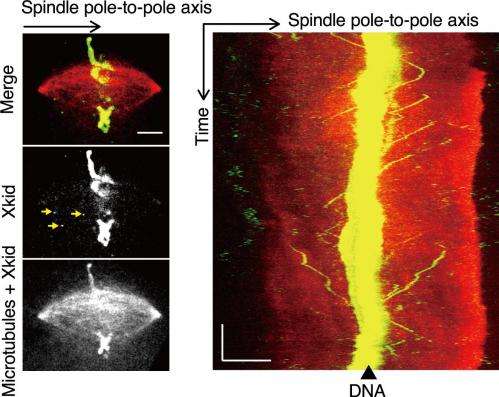
Spindles vary in size depending on the cell type. Those used in our study were self-organized spindles in Xenopus egg extracts, and they were relatively large, at 30 to 50 µm in length. This means that it is optically difficult to detect movements of fluorescence-labeled single molecular or ensembles of Xkid within a spindle under microscope. To overcome this, we used strongly fluorescent nanoparticles called quantum dots. By binding quantum dots to Xkid, we were able to determine the tracks of Xkid movements within such large spindles using fluorescence microscopy. Xkid molecules bound to quantum dots were located on chromosomes (DNA) as well as on microtubules (left; indicated by yellow arrows). Xkid present on microtubules traveled along the spindle pole-to-pole axis at 120 to 140 nm/sec (right). The scale bars for the horizontal and vertical axes indicate 10 µm and 100 sec, respectively.
Results and conclusions drawn from the study
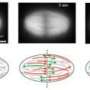
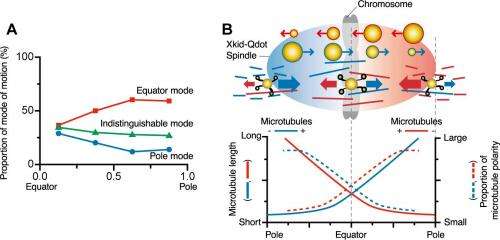
We found that Xkid traveled long distances (mean ≈5 µm; maximum 17 µm) along the oriented bundle of microtubules, moving from one microtubule to another along the way. We also observed Xkid moving mostly in the direction corresponding to the distribution of microtubule polarity, leading to the accumulation of Xkid near the spindle equator. This is consistent with the assembly and alignment of chromosomes near the spindle equator in metaphase. Our findings are the first molecular-level demonstration of the role of Xkid in directing chromosomes toward the spindle equator. Near the spindle poles, more Xkid molecules moved in the equator mode (toward the spindle equator), whereas near the spindle equator, all modes of motion were seen equally frequently (A). The distributions of polarity and length of microtubules were symmetrical with respect to the spindle equator, and corresponded with the predominant direction of Xkid movements (B). The size of the circles representing Xkid-Qdot (quantum dot) indicates the proportion of direction of movement in each location within the spindle.
Potentially universal effects and social significance of the study
When cells divide, segregation of chromosomes must take place with great precision. Uneven segregation of chromosomes can cause various forms of medical disorders including severe illnesses and malignant transformation of tumors. We determined the movement of a molecular motor involved in chromosome segregation within an intact spindle, a finding that has great medical implications. Furthermore, we showed that the oriented structure of microtubules within a spindle functions as a reaction field for the movement of Xkid, which is invaluable knowledge in exploring the mechanisms of materials transport in biological systems. One day we might be able to take control of chromosome segregation and materials transport by manipulating molecular motors and their reaction fields.
More information: Read the research paper: www.nature.com/srep/2013/13093 … /full/srep02808.html
Journal information: Scientific Reports
Provided by Waseda University