Researchers perform DNA computation in living cells
(Phys.org) —Chemists from North Carolina State University have performed a DNA-based logic-gate operation within a human cell. The research may pave the way to more complicated computations in live cells, as well as new methods of disease detection and treatment.
Logic gates are the means by which computers "compute," as sets of them are combined in different ways to enable computers to ultimately perform tasks like addition or subtraction. In DNA computing, these gates are created by combining different strands of DNA, rather than a series of transistors. However, thus far DNA computation events have typically taken place in a test tube, rather than in living cells.
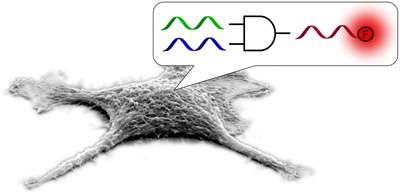
NC State chemist Alex Deiters and graduate student James Hemphill wanted to see if a DNA-based logic gate could detect the presence of specific microRNAs in human cells. The researchers utilized a DNA-based logic gate known as an "AND" gate that was engineered to respond to the presence of two specific microRNAs – known as miRNA-21 and miRNA-122.
Just as computer operations utilize different inputs to create a particular output, the researchers' DNA-based Boolean logic gate was activated only when both miRNA-21 and miRNA-122 "inputs" were present in cells. If they were present, the gate generated an "output" by releasing a fluorescent molecule.
Deiters believes that use of these logic gates could lead to more accurate tests and treatments for human disease, especially cancer.
"The fluorescent molecule we used in this logic-gate design could be useful as a marker that identifies a cancer cell," he says. "Or, instead of directing the gate to release a fluorescent molecule in the presence of particular microRNAs, we could attach therapeutic agents that are released to treat the disease itself."
Their results appear in the Journal of the American Chemical Society. The research was funded in part by grants from the American Chemical Society and the American Cancer Society.
More information: "DNA Computation in Mammalian Cells: microRNA Logic Operations" James Hemphill and Alexander Deiters, North Carolina State University, Journal of the American Chemical Society, 2013.
Abstract: DNA computation can utilize logic gates as modules to create molecular computers with biological inputs. Modular circuits that recognize nucleic acid inputs through strand hybridization activate computation cascades to produce controlled outputs. This allows for the construction of synthetic circuits that can be interfaced with cellular environments. We have engineered oligonucleotide AND gates to respond to specific microRNA (miRNA) inputs in live mammalian cells. Both single- and dual-sensing miRNA-based computation devices were synthesized for the cell-specific identification of endogenous miR-21 and miR-122. A logic gate response was observed with miRNA expression regulators, exhibiting molecular recognition of miRNA profile changes. Nucleic acid logic gates that are functional in a cellular environment and recognize endogenous inputs significantly expand the potential of DNA computation to monitor, image, and respond to cell-specific markers.
Journal information: Journal of the American Chemical Society
Provided by North Carolina State University




















