Molecular motors of nucleic acid: Researchers work to improve screening of helicase-targeting drugs
European scientists investigated the dynamic unfolding of DNA during replication by generating a tool that could subsequently be applied to screen helicase-targeting drugs for infection and oncologic applications.
In order to study the mechanical unfolding and refolding of various molecules including proteins and nucleic acids, and determine misfolded states, special equipment and techniques are required. To this end, optical tweezers and atomic force microscopes are proving extremely versatile tools that facilitate access to the inner functioning of biomolecules at an unprecedented level of detail.
The European Sminafel project focused on the activity of the helicase enzymes that assist the replication-repair of DNA. By hydrolysing adenosine triphosphate (ATP), these proteins convert chemical energy to the unzipping of the DNA double helix.
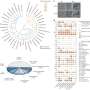
Scientists developed and optimised an optical tweezer-related technology that enabled the investigation of helicase function. More specifically, a DNA hairpin was fixed onto coated beads between a micropipette and an optical trap, and fluxing of the helicase and ATP solutions was facilitated through a microfluidics system. Various parameters of the system, including the valves and the length of the DNA molecule were standardised to allow efficient opening of the DNA hairpin, allowing the measurement of helicase activity.
Experimental results showed that the amplitude of fluctuations in the helicase activity remained constant independently of ATP concentration. The only determinant factors proved to be the opening-closing fluctuations of the replication fork.
The Sminafel technology constituted a significant step towards understanding the functioning of molecular motors involved in the DNA molecular repair and duplication machinery. The developed system is envisioned to provide a unique tool for studying various biomolecules in detail.
Provided by CORDIS