Biophysicists unravel secrets of genetic switch
When an invading bacterium or virus starts rummaging through the contents of a cell nucleus, using proteins like tiny hands to rearrange the host's DNA strands, it can alter the host's biological course. The invading proteins use specific binding, firmly grabbing onto particular sequences of DNA, to bend, kink and twist the DNA strands. The invaders also use non-specific binding to grasp any part of a DNA strand, but these seemingly random bonds are weak.
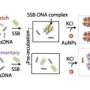
Emory University biophysicists have experimentally demonstrated, for the fist time, how the nonspecific binding of a protein known as the lambda repressor, or C1 protein, bends DNA and helps it close a loop that switches off virulence. The researchers also captured the first measurements of that compaction.
Their results, published in Physical Review E, support the idea that nonspecific binding is not so random after all, and plays a critical role in whether a pathogen remains dormant or turns virulent.
"Our findings are the first direct and quantitative determination of non-specific binding and compaction of DNA," says Laura Finzi, an Emory professor of biophysics whose lab led the study. "The data are relevant for the understanding of DNA physiology, and the dynamic characteristics of an on-off switch for the expression of genes."
C1 is the repressor protein of the lambda bacteriophage, a virus that infects the bacterial species E. coli, and a common laboratory model for the study of gene transcription.
The virus infects E. coli by injecting its DNA into the host cell. The viral DNA is then incorporated in the bacterium's chromosome. Shortly afterwards, binding of the C1 protein to specific sequences on the viral DNA induces the formation of a loop. As long as the loop is closed, the virus remains dormant. If the loop opens, however, the machinery of the bacteria gets hi-jacked: The virus switches off the bacteria's genes and switches on its own, turning virulent.
"The loop basically acts as a molecular switch, and is very stable during quiescence, yet it is highly sensitive to the external environment," Finzi says. "If the bacteria is starved or poisoned, for instance, the viral DNA receives a signal that it's time to get off the boat and spread to a new host, and the loop is opened. We wanted to understand how this C1-mediated, loop-based mechanism can be so stable during quiescence, and yet so responsive to switching to virulence when it receives the signal to do so."
Finzi runs one of a handful of physics labs using single-molecule techniques to study the mechanics of gene expression. In 2009, her lab proved the formation of the C1 loop. "We then analyzed the kinetics of loop formation and gained evidence that non-specific binding played a role," Finzi says. "We wanted to build on that work by precisely characterizing that role."
Emory undergraduate student Chandler Fountain led the experimental part of the study. He used magnetic tweezers, which can pull on DNA molecules labeled with miniscule magnetic beads, to stretch DNA in a microscope flow chamber. Gradually, the magnets are moved closer to the DNA, pulling it further, so the length of the DNA extension can be plotted against the applied force.
"You get a curve," Finzi explains. "It's not linear, because DNA is a spring. Then you put the same DNA in the presence of C1 protein and see how the curve changes. Now, you need more force to get to the same extension because the protein holds onto the DNA and bends it."
An analysis of the data suggests that, while the specific binding of the C1 protein forms the loop, the non-specific binding acts like a kind of zipper, facilitating the closure of the loop, and keeping it stable until the signal comes to open it.
"The zipper-like effect of the weaker binding sites also allows the genetic switch to be more responsive to the environment, providing small openings that allow it to breathe, in a sense," Finzi explains. "So the loop is never permanently closed."
The information about how the C1 genetic switch works may provide insights into the workings of other genetic switches.
"Single-molecule techniques have opened a new era in the mechanics of biological processes," Finzi says. "I hope this kind of experiment will lead to better understanding of how our own DNA is compacted into chromosomes, and how it unravels locally to become expressed."
Journal information: Physical Review E
Provided by Emory University