Predicting the microbial 'weather'
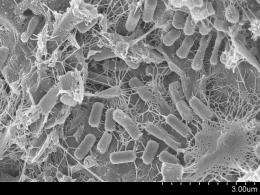
New computer models are letting scientists forecast changes in the population of microbes in the English Channel up to a week in advance.
Environmental microbiologist Jack Gilbert of the U.S. Department of Energy's Argonne National Laboratory heads the Earth Microbiome Project, an initiative to sample and analyze DNA from bacteria, viruses, algae and fungi across the world. Our environment is full of microbes that affect everything from human health to climate change, and these microbes are constantly in flux. One of the project’s goals is to develop models that can predict fluctuations in advance.
“Bacteria mediate everything: your health, your dog’s health, the crops in the field,” Gilbert said. “Without bacteria, there’d be no life. They’ve been around for 3.8 billion years. If they change, we change. We have to.”
Researchers collected years of data on the microbes that live in the English Channel. Each change reverberates up the food chain; one season’s abundance of particular algae can affect how much cod fisherman catch that year.
The team’s new computer models, which are much more accurate, can forecast microbial populations in the Channel a week ahead with about 95 percent accuracy.
The new versions sew together the interactions of many different viruses, bacteria and algae. “This is a radical change in the way we model,” Gilbert said.
Previous models didn’t work as well, Gilbert said, because they didn’t account for the hundreds of other species that affect the numbers of any one species of microbe.
“It’s as if you tried to predict the warthog population without also looking at the grasses and berries they eat, or the population of lions and crocodiles, which eat them, and the animals that affect them as well,” Gilbert explained.
The group is already collaborating with Argonne scientists who work on climate prediction. Biodiversity affects climate, because microbes and plants traffic in carbon dioxide—and thus help shape the future of the world’s climate. Adding the actions of microbes to the climate model will result in better predictions.
The next challenge planned for the Earth Microbiome Project is to build similar models for soil in the United States, and it’s harder. The English Channel was easier to model, Gilbert said, because water is easier to sample. When you scoop a cup of water out of the Channel to test it for microbes, the rest of the Channel mixes right back together. Taking a cup of dirt out of the soil creates a hole that doesn’t immediately get filled, which changes the microbial community—and that can affect the results when you return month after month to sample the same spot.
The study, “Predicting bacterial community assemblages using an artificial neural network approach,” was published in Nature Methods. Gilbert works with Argonne and the University of Chicago; other authors on the paper are Argonne’s Peter Larsen and Dawn Field at the U.K.’s Natural Environment Research Council. The research was supported by Argonne National Laboratory through Laboratory Directed Research and Development funding.
More information: Nature Methods (2012) doi:10.1038/nmeth.1975
Journal information: Nature Methods
Provided by Argonne National Laboratory


















