New imaging techniques reveal the workings of supramolecular nanomachines
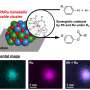
Supramolecules comprising many kinds of proteins and nucleic acids are present in all living organisms. Often with precise structures and a variety of parts, these supramolecules exhibit complex movements and exhaustive functions, essentially behaving like nanomachines. “How do these tiny supramolecular nanomachines work in the bodies of living organisms? I want to know the mechanism behind their actions,” says Koji Yonekura, associate chief scientist of the Biostructural Mechanism Laboratory in the Photon Science Research Division of the RIKEN SPring-8 Center. Because function is closely related to form, the action mechanism cannot be understood without clarifying the conformations of the components of supramolecules. Yonekura is working to develop new techniques such as cryo-electron microscopy to elucidate the action mechanism behind these supramolecular nanomachines.
The wonder of the flagellum
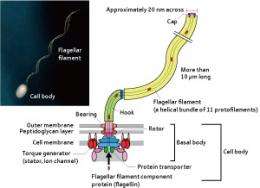
Escherichia coli and Salmonella enterica exhibit active movements, including moving towards places where nutrients are available and moving away from places of low temperature. The flagellum, an appendage that protrudes from bacterial surfaces, produces the driving force for these movements. The bacterial flagellum has a supramolecular structure of about 30 different proteins, comprising a basal body that works as a rotary motor, a flagellar filament that revolves like a propeller, and a hook that joins the two.
“As the ions flow into the flagellar cell, the rotor in the basal body revolves at up to 300 revolutions per second, so that the flagellar filament revolves like a propeller and causes the bacterium to move forward. The flagellum is like a well-designed nanomachine. I want to know how complex biological nanomachines like these work in living organisms,” says Yonekura. After learning the structural analysis of biomolecules by electron microscopy from Chikashi Toyoshima at the Tokyo Institute of Technology (currently at the Institute of Molecular and Cellular Biosciences, The University of Tokyo) in 1997, Yonekura joined the Namba Protonic Nanomachine Project (directed by Keiichi Namba, Graduate School of Frontier Biosciences, Osaka University) in an Exploratory Research for Advanced Technology (ERATO) program sponsored by the Japan Science and Technology Agency. Since then, he has been engaged in elucidating the mechanism behind the actions of the flagellum.
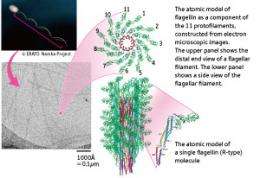
The flagellar filament is more than 10 μm in length, much longer than the main cell body of the bacterium, which measures just 1–2 μm. Each flagellar filament consists of a tubular bundle of 11 thin, long filaments known as protofilaments. “If the flagellar filament were linear, no propelling force would be generated even when it revolves at an extremely high speed. The reason why the bacterium can move its body forward is because the flagellar filament has a superhelical structure with a slight curvature. However, the very fact that the flagellar filament assumes this form is wonderful.”
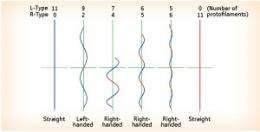
The flagellar filament is made up of a stack of flagellin, a protein produced in the cell body. When a form of protein of the same size and shape is stacked, only a linear structure is produced. Flagellin, however, occurs in structurally distinct left- and right-handed (L- and R-) types. As the L- and R-type protofilaments coexist in combination at different ratios, the flagellar filament can be transformed between the L and R structures . However, this is not the total mechanism behind bacterial movement by flagellar revolution. The bacterium repeats a motor pattern in which it goes forward for 2–3s, then tumbles and changes direction to propel itself forward again. What mechanism does the bacterium use to change its swimming direction?
It is known that when the bacterium changes direction, the counterclockwise rotor revolves in the reverse direction for 1 ms. With this motor reversal, a torsional force is exerted on the flagellar filament, causing some of the L-protofilaments to turn right-handed, and at this time the bacterium changes direction with the imbalance. This is the conjectured mechanism behind the directional switching in the bacterium. “The mechanical details of the directional change cannot be understood without clarifying the conformation of flagellin,” says Yonekura. However, this is a difficult task.
Cryo-electron microscopy and a new technique for helical reconstitution
A representative method for examining protein conformations is X-ray crystallography, in which a prepared protein crystal is exposed to an X-ray beam. Colliding with the crystal, X-rays are scattered by the electrons around the atomic nuclei, interfering with each other to produce a diffraction image. The distribution of electrons in the sample material is determined from the position and intensity of the diffraction to show how the atoms are arranged. A crystal is used because the orderly atomic arrangement ensures clear, regular diffraction images, which allow the investigator to examine the conformation at high resolution.
“However, X-ray crystallography cannot be applied to the flagellar filament because it is quite difficult to create a crystal of fibrous protein. Hence, we decided to use an electron microscope. In electron microscopy, the sample is exposed to an electron beam. The use of an electron microscope offers the advantage that a real image of the sample can be seen. However, the energy of electron beams is so intense that the sample is destroyed if it is treated as it is.”
To solve this problem, Yonekura used cryo-electron microscopy. “The sample is frozen rapidly and enclosed in ice. By keeping the sample under cooling conditions at –269 °C, the damage it experiences due to electron irradiation during observation is reduced dramatically. Additionally, because the procedure takes place while the sample is in ice, another advantage is the ability to obtain observations in nearly the same state as in a living organism in aqueous solution.”
However, cryo-electron microscopy alone does not enable conformational data to be obtained at high resolution because a high-intensity electron beam must be applied to obtain the desired level of resolution. “Even though protein damage is reduced markedly, it is still severe, so the exposure to the electron beam must be minimized. The images taken under these conditions contain a great deal of noise, and almost nothing is visible. To extract useful information from the noisy image at high resolution, a special technique is required for image analysis.”
Because of this, Yonekura has developed a technique for image analysis to reconstitute conformations using helical symmetry. “To obtain information at high resolution from noisy images, a common approach is to take and average out a large number of images. However, since it was thought that an astronomical number of images would be required to obtain data at sufficient resolution, the conventional approach was impractical. Using the new technique of helical reconstitution we have developed, the conformation of a sample can be visualized at a workable resolution by totaling about 100 micrographs, provided that the sample has a helical structure like the flagellar filament. When we analyze a sample that has a helical structure, molecules aligned in various directions are visualized in one photograph. We utilize them efficiently.”
The mechanism of flagellar actions now revealed
In 2003, Yonekura succeeded in analyzing the conformation of R-flagellin as a component of the flagellar filament using a combination of cryo-electron microscopy and the new technique of helical reconstitution. The resolution they achieved was close to 4 Å, or just 0.4 nm. Although that was a groundbreaking achievement that allowed the analyst to clarify the atomic arrangement in the amino acids that constitute flagellin, Yonekura and his colleagues continued to conduct their challenging work. “Analyzing the conformation of R-flagellin only allows us to know one state of the flagellar filament. The mechanism behind the morphological transition cannot be known without analyzing the conformation of the L-type.”
Then in 2008, Yonekura established his Biostructural Mechanism Laboratory at the RIKEN SPring-8 Center. At last, in March 2010, the laboratory succeeded in analyzing the conformation of the L-flagellin filament. The analysis took much time to develop because L-flagellin is unstable, but finally the molecular mechanism of the directional change in bacterial swimming was revealed. “By comparing the conformations of the L- and R-types, we find little change in the inner portion of the flagellar filament. On the other hand, the outer portion was found to change very flexibly. On motor reversal, part of the flagellin transits from the L-type to the R-type, switching the direction of the flagellar filament helix. As a result, the bacterium changes its swimming direction.”
The flagellar filament meets two contradictory requirements. One is the toughness needed to endure high-speed rotation, and the other is the flexibility required for the transition between the L- and R-types. “The conformation of flagellin really reconciles this toughness and flexibility. In fact, when we succeeded in analyzing the conformation, I was very impressed by the fact that the conformation is finely fabricated,” says Yonekura. “Actually, the conformation we determined proved to be rather different from the predicted structure. I realized again the importance of actually seeing what happens.
“In recent years, there has been a trend for research achievements to be influenced by the researchers’ ability to purchase expensive and sophisticated equipment. This is not very enjoyable. To make extensive efforts to allow myself to do what has not been done so far-that’s interesting. I would like to stick to this approach because a strong point of research is developing an ‘only-one’ technology.”
Three-dimensional crystallography of very fine crystals by electron microscopy
“I want to clarify the conformations of proteins at the highest possible resolution,” emphasizes Yonekura. He has devoted himself to developing a broad range of analytical techniques.
“A big feature of electron microscopy resides in dispensing with the need for sample crystallization. Despite this, crystals with a regular atomic arrangement are still attractive for those who want to examine the conformations of substances at high resolution. Even proteins that are usually unlikely to crystallize happen to form very fine crystals. We considered analyzing them by electron microscopy, but no suitable techniques were available. So we are developing a new technique in cooperation with Prof. Toyoshima at The University of Tokyo, Hitachi High-Technologies Corporation and Hitachi High-Tech Fielding Corporation.”
Electrons are 100,000 times more energetic than X-rays in terms of their potential for interactions with substances. This makes it possible to obtain information on atomic arrangement even in extremely small crystals. Recently, analytical techniques for very fine crystals using X-rays have been developed, and it is becoming possible to analyze crystals several micrometers long at SPring-8, the synchrotron radiation facility at the RIKEN Harima Institute. “Electron microscopy makes it possible to analyze even smaller crystals less than 1 μm in length. In membrane proteins and supramolecular complexes, which mediate a wide variety of biological activities and serve as drug discovery targets, only very fine crystals are formed in some cases, so this technique will be an important analytical tool.”
Crystallography using electron microscopy offers another major advantage. “While X-rays are scattered by electrons, electrons are influenced by electrical charge. As such, electron beams allow us to know not only how atoms are arranged in crystals, but also which portions of a protein are positively charged and which portions are negatively charged. When a protein is in action, the distribution of charge is very important.”
Yonekura targets the flagellar ion channel in his crystallographic work using electron microscopy. The ion channel is present in the basal body, through which ions enter the cell from outside. The ionic flow drives the rotor in the basal body to allow the flagellar filament to rotate. “Because ions are charged particles, it is possible to track their pathways. So far, no one in the world has been successful in three-dimensional crystallography of very fine crystals using electron microscopy. I hope that we will achieve this in a few years time.”
Another target of his is the telomere, a structure at the end of the chromosome. The telomere is said to shorten upon each cell division and control cell longevity. Conversely, the telomere elongates in cancer cells. “The telomere is a large, complex supramolecule comprising DNA and proteins. I want to unveil the mechanism that controls the length of the telomere by determining its conformation.”
Making the best use of all technical resources
In 2009, Yonekura and Saori Maki-Yonekura, a researcher in the Protein Crystallography Research Group at the RIKEN SPring-8 Center, were awarded the 2009 Ernst Ruska Award for their work entitled ‘Contribution toward elucidating the mechanisms of biological macromolecular machines by cryo-electron microscopy’. The award was established in 1980 by the German Society for Electron Microscopy in commemoration of Ernst Ruska, 1986 winner of the Nobel Prize in Physics and the inventor of electron microscopy. “I hear that the award is bestowed for achievements that involve a technical innovation, as well as a groundbreaking application of electron microscopy to visualize something previously invisible. This is what I have been aiming at, and I am very proud to have received the award.”
Yonekura is looking forward to utilizing the X-ray Free Electron Laser (XFEL) facility under construction at the RIKEN Harima Institute, which is scheduled to be completed in fiscal 2011. “The XFEL is expected to enable us to perform conformational analyses on whole cells and organelles as they are. We are now developing a new technique for analyzing biological samples by a combination of the XFEL and cryo-electron microscopy, jointly with Prof. Masayoshi Nakasako in the Department of Physics of the Faculty of Science and Technology at Keio University, as well as Director Masaki Yamamoto of the Research Infrastructure Group in the Advanced Photon Technology Division of the RIKEN SPring-8 Center and others. I am aiming at elucidating the mechanism behind the actions of nanomachines in living organisms. To this end, I am not fixed only on electron microscopy. Here at the RIKEN Harima Institute, a radiation facility with the world’s highest performance is in operation, and the associated equipment is available in a comprehensive link-up. I want to clarify the mechanism behind the actions of nanomachines by making the best use of the technical resources that have been compiled here over a long period. Many people are working to develop a broad range of equipment, and I believe that I can create a new technique by listening to their valuable advice and combining it with my own knowledge. I am now enjoying my research a lot.”
Provided by RIKEN