Greenhouse gas-producing enzyme may yield insights into earliest oxygen-breathing ancestor evolution
Every year, nitrogen-metabolizing bacteria in the soil and seas churn out more than ten billion kilograms of nitrous oxide (N2O) gas as they respire in these oxygen-deficient environments.
Nitric oxide reductase (NOR) enzymes are the powerhouse underlying production of this gas, taking pairs of nitric oxide (NO) molecules and transforming them into N2O and water via a chemical reaction known as ‘reduction’. These enzymes also help pathogenic bacteria to evade destruction by the immune system, as some T cells use NO as a chemical weapon against infectious agents.
More generally, scientists are interested in the potential to employ these enzymes as a tool for synthesizing useful, customized molecules for a variety of applications. “The nitrogen-oxygen bond cleavage and nitrogen–nitrogen bond formation reactions executed by these enzymes are the essence of chemistry,” says Yoshitsugu Shiro of the RIKEN SPring-8 Center in Harima.
Although a great deal is known about the biochemical properties of these proteins, scientists have found it challenging to determine the structure of bacterial NOR in fine detail. Now, after seven years of hard work, Shiro and colleagues have finally obtained the first such structure for NOR from Pseudomonas aeruginosa, a pathogenic bacterium associated with opportunistic infections in immune-compromised patients (Fig. 1).
Not-so-distant relations
Although all NOR enzymes execute essentially the same chemical reaction to produce N2O, they can be subdivided into three major classes: cNOR, qNOR and qCuNOR. The enzyme crystallized by Shiro and colleagues (Fig. 2) belongs to the cNOR family, and contains a subunit known as cytochrome c that also enables bacteria to engage in aerobic (oxygen-driven) respiration. This ability to switch from oxygen-based to nitrogen-based respiration is highly beneficial for survival in the oxygen-poor conditions deep in the soil or beneath the waves.
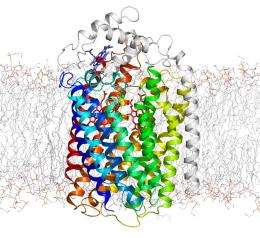
Accordingly, cNORs are thought to be closely related to the cytochrome oxidases (COX), enzymes that play a central role in aerobic respiration, and these new findings have revealed a number of structural parallels between the two. “There is a long history of research into these respiratory enzymes, COX and NOR, and a lot of knowledge on NOR has been accumulated by biochemical, chemical, molecular biology and microbiological studies,” says Shiro. “From these points of view, our NOR structure is not surprising, but seeing is believing!”
The reduction process is dependent on the directional transport of electrons and protons, and COX and cNOR appear to closely resemble one another in terms of the structure of their electron-transfer networks. Both enzymes depend on precisely positioned metal ions to enable electron transport, and the four iron atoms contained within cNOR are arranged in a configuration that closely resembles COX, maintained via interactions between these positively charged iron atoms and a set of evolutionarily conserved, negatively charged histidine and glutamate amino acids.
On the other hand, the researchers observed some notable differences with regard to the movement of protons. Both COX and cNOR are bound within membranes, but COX contains channels that are believed to direct the flow of protons across the membrane from the interior of the cell. This flow helps generate electrical potential that subsequently powers a variety of cellular motors. However, cNOR lacks such membrane-spanning channels, and protons entering the enzyme from the exterior of the cell only make it as far as the membrane interior, where the reductase catalytic site is located.
Back to the beginning
Even with a structure in hand for this well-studied enzyme, a number of mysteries remain to be addressed. For example, these data are insufficient to resolve an ongoing debate over the fine details of the N2O production mechanism. Shiro and colleagues were readily able to identify the two iron atoms involved in catalysis, but their structure reveals insufficient space at this ‘active site’ to accommodate the two molecules of NO believed to be required for this reaction.
“The NOR active site is tightly packed and very crowded,” says Shiro. “This observation suggests that some conformational change [is] needed to achieve catalytic turnover, but no one knows of any such conformational change so far.” Resolving this issue will require the acquisition of additional, high-resolution structures that might offer clear snapshots of the enzyme at intermediate stages in the catalytic process.
This structure offers tentative support for the hypothesis that the COX aerobic respiratory machinery originally evolved from NOR enzymes, although additional work will clearly be required to confirm this. Unlike NOR, which exclusively employs iron ions, COX makes use of both copper and iron for catalysis, and the researchers have tentatively identified a few amino acid changes that might have enabled this transition to take place. In addition, although NOR lacks the ‘K-channel’ that allows COX to deliver protons from the cytoplasm, Shiro’s team has identified some structural elements that could potentially represent early evolutionary precursors in the formation of this channel.
In future studies, Shiro plans to develop experimental tests for some of these still-speculative models. “We want to follow the molecular evolution of the respiratory enzymes from anaerobic to aerobic conditions on Earth, from NOR to COX,” he says. “Using mutagenesis, based on our structural comparisons, we are hoping to convert NO-reducing NOR into oxygen-reducing COX.” For the present, though, he is optimistic that this structure will give a boost to researchers seeking to understand and manipulate this enzymatic process. “Scientists worldwide who are interested in NO reduction can enter a new stage of NOR research with this structure,” he says.
More information: Hino, T., et al. Structural basis of biological N2O generation by bacterial nitric oxide reductase. Science 330, 1666–1670 (2010). www.sciencemag.org/content/330/6011/1666.abstract
Provided by RIKEN















