3-D tracking of single molecules inside cells
Researchers at the University of Texas Southwestern Medical Center and the University of Texas at Dallas are reporting today at the 55th Annual Biophysical Society Annual Meeting in Baltimore, MD how they are using a novel 3D cell imaging method for studying the complex spatial-temporal dynamics of protein transport, providing a solution to this fundamental problem in cell biology.
According to the authors of the study, imaging such highly dynamic processes in the cell and in 3D poses major technical challenges in a complex cell monolayer due to cell-to-cell variations in thickness and temporal properties of protein transport. Previous imaging techniques were slow and suffered from poor z-localization/3D-tracking capability.
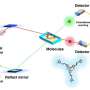
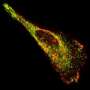
Using a combination of multifocal plane microscopy (MUM) and nanodot labeling technology, the researchers were able to label single molecules in live cells and track their movement and their interaction with other molecules in a thick cell sample for extended periods of time.
Sripad Ram, the lead author of the study, explains that the main reason he and his colleagues developed these imaging techniques is to track the movement of therapeutic antibodies, which are engineered in their lab. "We want to know where these antibodies go and what they do once they enter the body," says Ram.
He adds that "current microscopy technologies are limited in that you can only image a single focal plane at any given time. If you want to image in three dimensions, you can only do so sequentially, but you end up imaging at the wrong place and at the wrong time thereby missing events over time….what we needed was a technology that could simultaneously image a sample across multiple planes, and that is what multifocal plane microscopy is all about."
More information: The presentation, "3D SINGLE MOLECULE TRACKING IN THICK CELLULAR SAMPLES USING MULTIFOCAL PLANE MICROSCOPY" by Sripad Ram, E. Sally Ward, and Raimund J. Ober is at 11:00 am. on Tuesday, March 8, 2011 in Ballroom IV of the Baltimore Convention Center. ABSTRACT: tinyurl.com/4zzt6ys
Provided by American Institute of Physics