Releasing the brakes
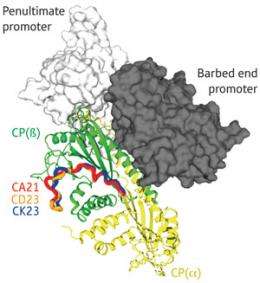
Two regulators of protein filament assembly use dramatically different -- and competing -- methods to inhibit a common target.
Actin-based protein filaments participate in biological activities ranging from cell migration to muscle contraction. These filaments can be highly dynamic, with individual actin molecules spontaneously attaching to or dissociating from the ends of the fiber. Typically, however, such activity is closely regulated by factors like actin capping protein (CP).
Filaments exhibit physical polarity, with extension specifically occurring at the ‘barbed’ end, and CP inhibits addition of new actin molecules by firmly seating itself at this end. CP is widely conserved in species ranging from yeast to humans and acts a crucial regulator for a variety of actin-mediated cellular functions.
Accordingly, cells also produce factors that help remove CP from filament ends, such as the V-1 and CARMIL proteins. Yasushi Nitanai at the RIKEN SPring-8 Center in Harima recently partnered with Nagoya University researchers Shuichi Takeda and Yuichiro Maeda to characterize the mechanisms employed by these two CP regulators via structural analysis1.
CP is composed of an α and a β subunit, each of which has a projecting ‘tentacle’ domain. Previous work from Takeda and Maeda showed that CP relies on the α tentacle to latch onto actin while the β tentacle stabilizes the complex2. Their work with Nitanai has now demonstrated that V-1 acts as a direct counter to this process, binding the same portions of the α tentacle that mediate actin binding and thereby physically preventing them from associating with the filament.
Takeda and colleagues identified a markedly different mechanism for CARMIL, based on data that revealed a surprisingly dynamic structure for CP. “We had believed that CP was a rigid molecule, and never imagined that it was an intrinsically flexible molecule, continuously undergoing twisting motions,” says Takeda. CARMIL appears to actively exploit this flexibility, interacting with CP via a relatively unstructured domain. This association does not physically obstruct actin binding, but instead constrains CP into an arrangement that reduces its affinity for both the barbed end of actin filaments and the V-1 inhibitor.
The team’s results are in keeping with previous findings indicating that CARMIL can bind to CP that is already bound to filament ends and triggers its rapid dissociation. “We were impressed with the way that CARMIL utilizes the intrinsic fluctuation of CP to suppress capping activity,” says Takeda. In future studies, he and his colleagues hope to apply alternative structural biology techniques, such as nuclear magnetic resonance, to better capture the subtle details of the dynamic interactions between CARMIL, V-1 and CP.
More information:
1.Takeda, S., Minakata, S., Koike, R., Kawahata, I., Narita, A., Kitazawa, M., Ota, M., Yamakuni, T., Maeda, Y. & Nitanai, Y. Two distinct mechanisms for actin capping protein regulation—steric and allosteric regulation. PLoS Biology 8, e1000416 (2010).
2.Narita, A., Takeda, S., Yamashita, A. & Maeda, Y. Structural basis of actin filament capping at the barbed-end: a cryo-electron microscopy study. The EMBO Journal 25, 5626-5633 (2006). www.nature.com/emboj/journal/v … 23/abs/7601395a.html
Provided by RIKEN